
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,084,589 – 13,084,693 |
| Length | 104 |
| Max. P | 0.555785 |

| Location | 13,084,589 – 13,084,693 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 79.89 |
| Mean single sequence MFE | -24.68 |
| Consensus MFE | -15.42 |
| Energy contribution | -15.53 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13084589 104 + 22407834 AGAGGCCAGU-AUGAGAAAAUA-----GCAAAAGAA----ACG---ACUGCAAAUUGUAUGGCAU-UAAGAGACACUGU-UGCCAGAUCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA ........((-((((...((((-----(((......----...---..))).........(((((-((((((.((((..-...........)))))))))))))))))))..)))))). ( -20.42) >DroPse_CAF1 85098 113 + 1 AGAUGUAGCG-UCGUGCACAUUCGUAUGUGUAAGAAAACCCCG---ACUGCAAAUUGUAUGGCAU-UAAGAGACAUUGU-UGCCAUAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA .........(-((((((((((....))))))).........))---)).((((((((...(((((-((((((.((((..-.((....))..)))))))))))))))...))))).))). ( -30.20) >DroEre_CAF1 66986 104 + 1 AGAGGCAAGU-UUGAGAAAAUA-----GUAGAAGAA----GCC---GCUGCAAAUUGUACGGCAU-UCAGAGACACUGU-UGCCAGAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA ....(((...-((((..(((((-----.((((((..----(((---(.(((.....)))))))..-.......((((..-.((....))..)))))))))).))))).))))...))). ( -24.60) >DroYak_CAF1 66208 105 + 1 AGAGGCAAGU-AUGUGAAAAUA-----GCUGAAGAA----ACG---ACUGCAAAUUGUAUGGCAU-UAAGAGACUCUGUUUGCCAGAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA .......(((-.(((....)))-----)))......----...---...((((((((...(((((-(((((((((..((((....))))..))).)))))))))))...))))).))). ( -22.90) >DroAna_CAF1 52708 109 + 1 ACGGACGGCUAACAAAAAAAGA-----AAAGAAGAA----ACGCCAACUGCAAAUUGUAUAGCAUUUAAGAGACAUUGU-UGCCAUAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA ......(((.............-----.........----..)))....((((((((...((((((.(((((.((((..-.((....))..)))))))))))))))...))))).))). ( -19.76) >DroPer_CAF1 81895 113 + 1 AGAUGUAGCG-UCGUGCACAUUCGUAUGUGUAAGAAAACCCCG---ACUGCAAAUUGUAUGGCAU-UAAGAGACAUUGU-UGCCAUAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA .........(-((((((((((....))))))).........))---)).((((((((...(((((-((((((.((((..-.((....))..)))))))))))))))...))))).))). ( -30.20) >consensus AGAGGCAACU_ACGAGAAAAUA_____GUAGAAGAA____ACG___ACUGCAAAUUGUAUGGCAU_UAAGAGACACUGU_UGCCAGAGCUAAGUGCUUUUAAUGUUUAUUAAUUAUGCA .................................................((((((((...(((((.((((((.((((....((....))..)))))))))))))))...))))).))). (-15.42 = -15.53 + 0.11)

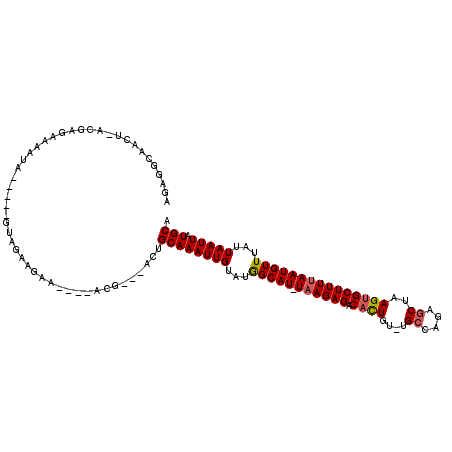
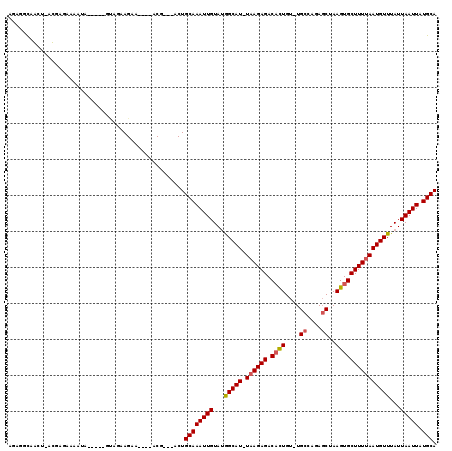
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:04 2006