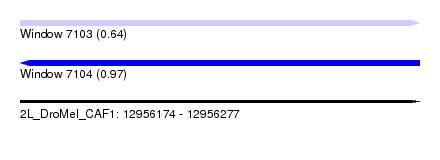

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,956,174 – 12,956,277 |
| Length | 103 |
| Max. P | 0.965062 |

| Location | 12,956,174 – 12,956,277 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 94.74 |
| Mean single sequence MFE | -32.75 |
| Consensus MFE | -29.70 |
| Energy contribution | -30.08 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644821 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12956174 103 + 22407834 CUCACUUGCUCUGCUCAACUGAUUGCUUACAGUGAGUGCAAACAGUAUUCUGAUUGGUUCAGGGCAAUCAGUGGCUGGGGAUCGCAGAACUACAGCGGGAUUU--- ....((((((((((...(((.((((....)))).))).....((((...(((((((.(....).)))))))..))))......)))))......)))))....--- ( -29.80) >DroSec_CAF1 936 103 + 1 CUCGCUUGCUCUGCUCAACUGAUUGCUUACAGUGAGUGCAAACAGUAUUCUGAUUGGUUCAGGGCAAUCAGUGGCUGGGGAUCGCAGAACUACAGCGGGAUUU--- ((((((.(.(((((...(((.((((....)))).))).....((((...(((((((.(....).)))))))..))))......))))).)...))))))....--- ( -34.50) >DroSim_CAF1 1039 105 + 1 CUCGCUUGCUCUGCUCAACUGAUUGCUUACACUGAGUGCAAACAGUAUUCUGAUUGGUUCAG-GCAAUCAGUGGCUGGGGAUCGCAGAUCUACAGCGGGAUUUUAA ((((((..(((.(((..(((((((((((..(((((((((.....)))))).....)))..))-)))))))))))).)))((((...))))...))))))....... ( -35.50) >DroYak_CAF1 1078 103 + 1 CUCGCUUGCUCUGCUCAACUGAUUGCUUACAGUGAGUGCAAACAGUAUUCUGAUUGGUUCAGGGCAAUCAGUGGCUGGAGAUCCCAGAACUACAUCGAGAUUU--- ((((.((((((((..(.(((.((((....)))).))).(((.(((....))).))))..))))))))...((((((((.....))))..))))..))))....--- ( -31.20) >consensus CUCGCUUGCUCUGCUCAACUGAUUGCUUACAGUGAGUGCAAACAGUAUUCUGAUUGGUUCAGGGCAAUCAGUGGCUGGGGAUCGCAGAACUACAGCGGGAUUU___ ((((((.(.(((((((.(((((((((((.....((((((.....))))))(((.....)))))))))))))).....)))....)))).)...))))))....... (-29.70 = -30.08 + 0.37)

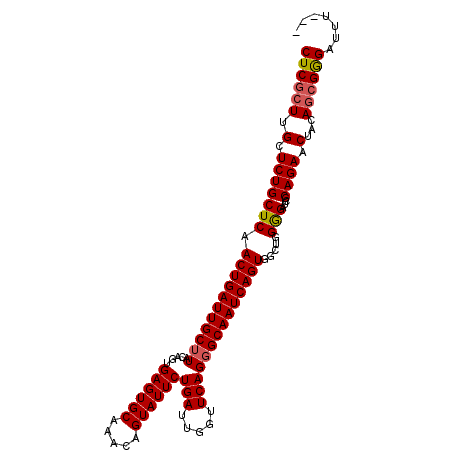
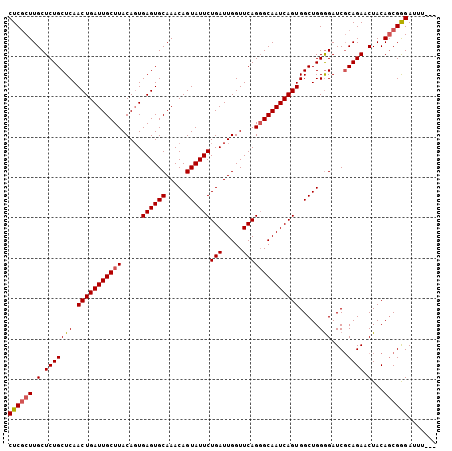
| Location | 12,956,174 – 12,956,277 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 94.74 |
| Mean single sequence MFE | -31.65 |
| Consensus MFE | -28.44 |
| Energy contribution | -29.00 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965062 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12956174 103 - 22407834 ---AAAUCCCGCUGUAGUUCUGCGAUCCCCAGCCACUGAUUGCCCUGAACCAAUCAGAAUACUGUUUGCACUCACUGUAAGCAAUCAGUUGAGCAGAGCAAGUGAG ---......((((...(((((((.........(.(((((((((.((((.....))))........(((((.....)))))))))))))).).))))))).)))).. ( -30.80) >DroSec_CAF1 936 103 - 1 ---AAAUCCCGCUGUAGUUCUGCGAUCCCCAGCCACUGAUUGCCCUGAACCAAUCAGAAUACUGUUUGCACUCACUGUAAGCAAUCAGUUGAGCAGAGCAAGCGAG ---......((((...(((((((.........(.(((((((((.((((.....))))........(((((.....)))))))))))))).).))))))).)))).. ( -32.70) >DroSim_CAF1 1039 105 - 1 UUAAAAUCCCGCUGUAGAUCUGCGAUCCCCAGCCACUGAUUGC-CUGAACCAAUCAGAAUACUGUUUGCACUCAGUGUAAGCAAUCAGUUGAGCAGAGCAAGCGAG .........((((.....(((((.........(.(((((((((-((((.....)))).((((((........))))))..))))))))).).)))))...)))).. ( -33.00) >DroYak_CAF1 1078 103 - 1 ---AAAUCUCGAUGUAGUUCUGGGAUCUCCAGCCACUGAUUGCCCUGAACCAAUCAGAAUACUGUUUGCACUCACUGUAAGCAAUCAGUUGAGCAGAGCAAGCGAG ---....((((.....((((((((....))..(.(((((((((.((((.....))))........(((((.....)))))))))))))).)..))))))...)))) ( -30.10) >consensus ___AAAUCCCGCUGUAGUUCUGCGAUCCCCAGCCACUGAUUGCCCUGAACCAAUCAGAAUACUGUUUGCACUCACUGUAAGCAAUCAGUUGAGCAGAGCAAGCGAG .........((((...(((((((.........(.(((((((((.((((.....))))........(((((.....)))))))))))))).).))))))).)))).. (-28.44 = -29.00 + 0.56)

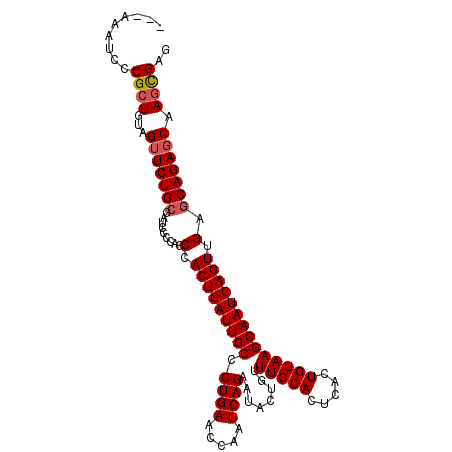
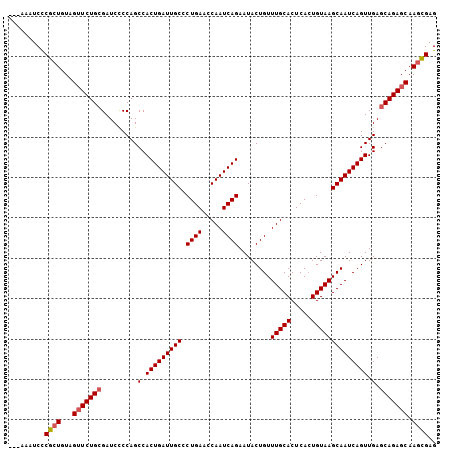
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:43 2006