
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,953,074 – 12,953,164 |
| Length | 90 |
| Max. P | 0.805860 |

| Location | 12,953,074 – 12,953,164 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 76.03 |
| Mean single sequence MFE | -16.15 |
| Consensus MFE | -10.30 |
| Energy contribution | -11.42 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.805860 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
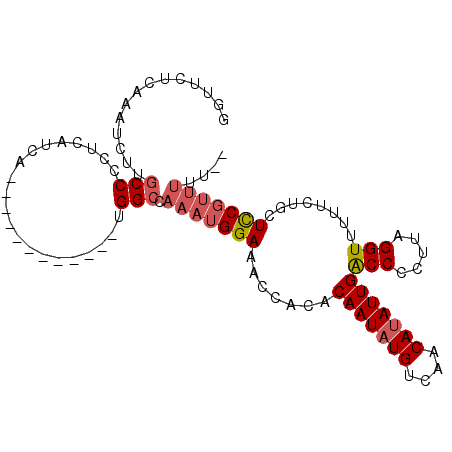
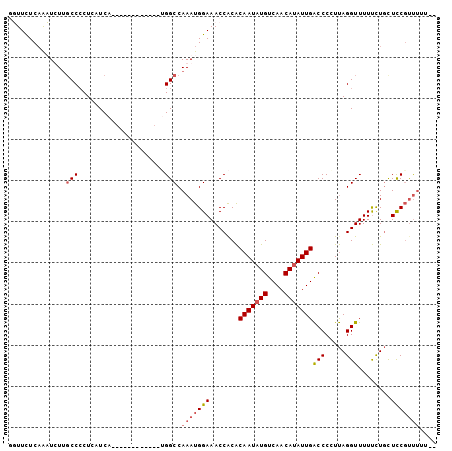
>2L_DroMel_CAF1 12953074 90 - 22407834 UGUUUUCUAUUCUUGCCACUUAUCA------------CGGCCAAAUGGAAACCACACAAUAUGUCAACAUAUUGACCCCUUAGGUUUUUUUGCUUCGUUUUC-- ((.((((((((...(((........------------.)))..)))))))).))..(((((((....)))))))(((.....))).................-- ( -16.50) >DroSec_CAF1 124887 92 - 1 GUUUCUCAAAUUUUGCCCCUCAUCA------------UGGCCAAAUGGAAACCACACAAUAUGUCAACAUAUUGACCCCUUAGGUUUUUCUGCUCCGUUUUUUU ((((((...((((.(((........------------.))).))))))))))....(((((((....)))))))(((.....)))................... ( -15.40) >DroSim_CAF1 130584 91 - 1 GUUUCUCAAAUUUUGCCCCUAAUCA------------GGGCCAAAUGGAAACCGCACAAUAUGUCAACAUAUUGACCCCUUAGGUUUUUCUGUUCCGUUUUUU- ..............((((.......------------)))).((((((((......(((((((....)))))))(((.....))).......))))))))...- ( -20.30) >DroYak_CAF1 127365 99 - 1 UGUUCUCAGUUCUGCCCCCUCAUCACACCUCAUUCCAUGGCCAAAUGGAAACCACGCAAUAUGUCAACACAUUGGCCCAUCAGGUUUUCCUCCUCCUCC----- ..........................((((..((((((......)))))).....((...((((....))))..)).....))))..............----- ( -12.40) >consensus GGUUCUCAAAUCUUGCCCCUCAUCA____________UGGCCAAAUGGAAACCACACAAUAUGUCAACAUAUUGACCCCUUAGGUUUUUCUGCUCCGUUUUU__ ..............(((.....................))).(((((((.......(((((((....)))))))(((.....)))........))))))).... (-10.30 = -11.42 + 1.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:39 2006