
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,945,232 – 12,945,337 |
| Length | 105 |
| Max. P | 0.898731 |

| Location | 12,945,232 – 12,945,337 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -31.58 |
| Consensus MFE | -20.98 |
| Energy contribution | -21.74 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.716032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12945232 105 + 22407834 AAAGGGAGGUGCAGCAGGGGAUCAAAAGCAACAAAAUUAACACAAACUAUUGCUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUUU------------GCUGCUGCUGAUGG ..........(((((((.(((.....(((((..................)))))(((..((((((((.........)))))))).))))))------------.)))))))...... ( -27.87) >DroSec_CAF1 117112 105 + 1 AAAGGGAGGAGCAGCAGGGGAUCAAAAGCAACAAAAUUAACACAAACUAUUGUUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUCU------------GCUGCUGCUGAUGG ......((.((((((((((........(((((((...............))))))).((((((((((.........)))))))).))))))------------)))))).))..... ( -37.16) >DroSim_CAF1 122334 102 + 1 AAAGGGAGGAGCAGCAGGGGAUCAAAAGCAACAAAAUUAACACAAACUAUUGUUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUCU------------GCU---GCUGAUGG .........((((((((((........(((((((...............))))))).((((((((((.........)))))))).))))))------------)))---)))..... ( -34.06) >DroEre_CAF1 124124 105 + 1 AAUGGGAGAAGCAUCCGGGAAUCAAAAGCAAGAAAAAUAACACAAACUUUUGUUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUCC------------GCUGCUGCUGAUGG .((.((...((((..((((........((((.((((...........)))).)))).((((((((((.........)))))))).)).)))------------).)))).)).)).. ( -24.90) >DroYak_CAF1 119428 117 + 1 AAUGGGAGGAGCAUUCGGGAAUCAAAAGCAACAAAAUUAACACAAACUUUUGUUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUCUGAUGCUGCUGGUGCUGCUGCUGAUGG ...((.((.(((((((((..((((...(((((((((...........))))))))).((((((((((.........)))))))).))...)))).)))..)))))).)).))..... ( -33.90) >consensus AAAGGGAGGAGCAGCAGGGGAUCAAAAGCAACAAAAUUAACACAAACUAUUGUUGCACACCACAAUGUAUCUAAAUUAUUGUGGCUGCUCU____________GCUGCUGCUGAUGG .......((((((((............(((((((...............)))))))....(((((((.........)))))))))))))))............((....))...... (-20.98 = -21.74 + 0.76)

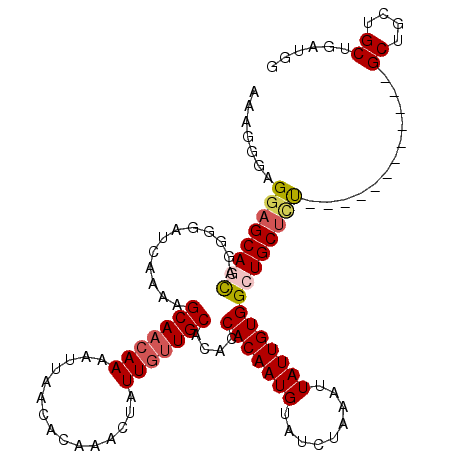
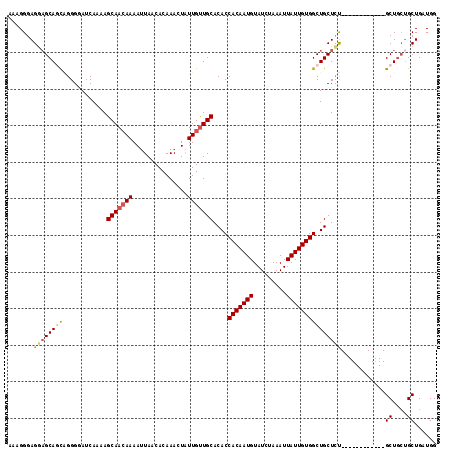
| Location | 12,945,232 – 12,945,337 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -30.16 |
| Consensus MFE | -21.12 |
| Energy contribution | -21.76 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898731 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12945232 105 - 22407834 CCAUCAGCAGCAGC------------AAAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAGCAAUAGUUUGUGUUAAUUUUGUUGCUUUUGAUCCCCUGCUGCACCUCCCUUU ......(((((((.------------...(((((((((((...........)))))))).)))((((((((..(......)..)))))))).........))))))).......... ( -32.10) >DroSec_CAF1 117112 105 - 1 CCAUCAGCAGCAGC------------AGAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAACAAUAGUUUGUGUUAAUUUUGUUGCUUUUGAUCCCCUGCUGCUCCUCCCUUU .....((((((((.------------.((.((((((((((...........))))))))..(((((((.((((......)))))))))))...)).))..))))))))......... ( -32.10) >DroSim_CAF1 122334 102 - 1 CCAUCAGC---AGC------------AGAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAACAAUAGUUUGUGUUAAUUUUGUUGCUUUUGAUCCCCUGCUGCUCCUCCCUUU .....(((---(((------------((.(((.(((((((...........))))))))))(((((((.((((......)))))))))))..........))))))))......... ( -31.10) >DroEre_CAF1 124124 105 - 1 CCAUCAGCAGCAGC------------GGAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAACAAAAGUUUGUGUUAUUUUUCUUGCUUUUGAUUCCCGGAUGCUUCUCCCAUU .....((.((((.(------------((.(((((((((((...........)))))))).)))..(((((((..(..........)..)))))))....)))..)))).))...... ( -27.20) >DroYak_CAF1 119428 117 - 1 CCAUCAGCAGCAGCACCAGCAGCAUCAGAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAACAAAAGUUUGUGUUAAUUUUGUUGCUUUUGAUUCCCGAAUGCUCCUCCCAUU .....((((((.((....)).))(((((((((.(((((((...........))))))))))(((((((((...........))))))))).)))))).......))))......... ( -28.30) >consensus CCAUCAGCAGCAGC____________AGAGCAGCCACAAUAAUUUAGAUACAUUGUGGUGUGCAACAAUAGUUUGUGUUAAUUUUGUUGCUUUUGAUCCCCUGCUGCUCCUCCCUUU ..(((((.((((((...............(((((((((((...........)))))))).))).((((....)))).........)))))).))))).................... (-21.12 = -21.76 + 0.64)

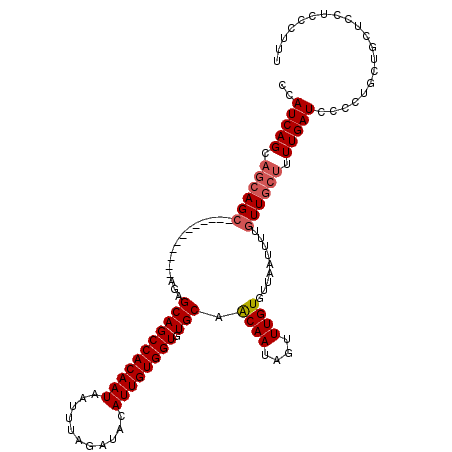
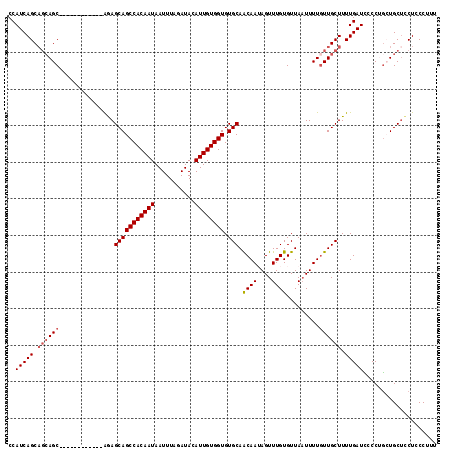
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:36 2006