
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,920,175 – 12,920,286 |
| Length | 111 |
| Max. P | 0.633476 |

| Location | 12,920,175 – 12,920,286 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.67 |
| Mean single sequence MFE | -31.27 |
| Consensus MFE | -19.46 |
| Energy contribution | -20.52 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.633476 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12920175 111 + 22407834 UAUGUCGUCGAGAAGAUUAAAGUCGUGGGCUGCACUUAUAUGGCAGCUUGUGGACUUGAUUUUACCCUGGCUAAGUCAAAAUUUGGAAGCAGAACACA---------UGCUUCUUACUCA .........(((((((((((..(((..((((((.(......)))))))..)))..))))))))....((((...))))......(((((((.......---------)))))))..))). ( -32.80) >DroSec_CAF1 99272 111 + 1 UAUGUCGUAGAAAAGAUUAAAGUCGUGGGCUGCACUUAUAUGGCAGCUUGCGGACUUGAUUUUACCCUGGCCAAGUCCAAAUUUGGAAGCAGAACACA---------UGCUUCUUACUCA ......(((....(((((......(..((((((.(......)))))))..)(((((((.............))))))).)))))(((((((.......---------))))))))))... ( -34.02) >DroSim_CAF1 103266 110 + 1 UAUGUCGUCGAGAAGAU-AAAGUCGUGGGCUGCACUUAUAUGGCAGCUUGUGGACUUGAUUUUACCCUGGCCAAGUCAAAAUUUGGUAGCAGAACACA---------UGCUUCUUACUCA .........(((((((.-.((((((..((((((.(......)))))))..).)))))............(((((((....)))))))((((.......---------)))))))).))). ( -31.80) >DroEre_CAF1 105949 120 + 1 UAUGUGGUCGAGAAGAUCAAAGUCGUUGGCUGCACCUACAUGGCGGCUUGUGGACUUGAUUUUACCCUGGCCAAGUCAAAAUUUGGCAGCAAAGUACAGGUUACUAAUGCCUCUUUCUCA ..((.((..(((((((((((.(((...((((((.........))))))....)))))))))))..(((((((((((....))))))).((...)).))))..........)))..)).)) ( -32.40) >DroYak_CAF1 101415 120 + 1 UAUGUAGUCGAGAAGAUCAAAGUCGUGGGCUGCACUUAUAUGGCAGCUUGUGGACUUGACUACACCCUGGCUAAGUCGAAAUUUGGAAGCAAAACACAGGUUACUACUUCCUCUUUCACA ..(((((((((.....)).((((((..((((((.(......)))))))..).))))))))))))..........((.((((...(((((...(((....)))....)))))..)))))). ( -35.50) >DroAna_CAF1 120014 100 + 1 UACGUGGUUGAGAAGAUCAAGGUGGUUGGCUGCACUUACAUGGCUGCCUGUGGACUAGAUUUUAAUAUUGCCUCGUUA------GGGAGUAAAGCAAAAAUGAACA-------------- ..(((..(((..((((((.((......(((.((.(......))).)))......)).))))))..((((.(((....)------)).))))...)))..)))....-------------- ( -21.10) >consensus UAUGUCGUCGAGAAGAUCAAAGUCGUGGGCUGCACUUAUAUGGCAGCUUGUGGACUUGAUUUUACCCUGGCCAAGUCAAAAUUUGGAAGCAAAACACA_________UGCUUCUUACUCA ............((((((((..(((..((((((.(......)))))))..)))..)))))))).......((((((....)))))).................................. (-19.46 = -20.52 + 1.06)

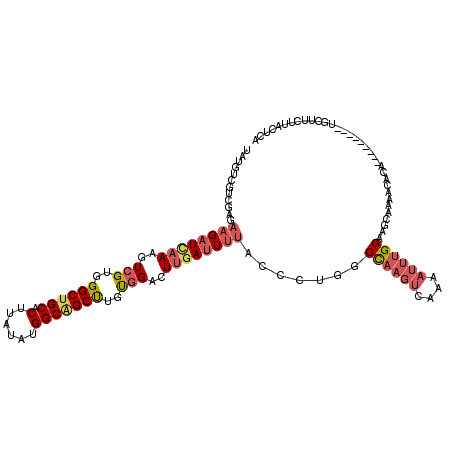
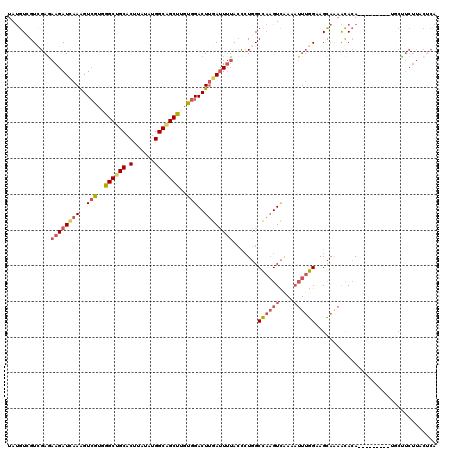
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:26 2006