
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,804,183 – 12,804,281 |
| Length | 98 |
| Max. P | 0.602441 |

| Location | 12,804,183 – 12,804,281 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.59 |
| Mean single sequence MFE | -27.73 |
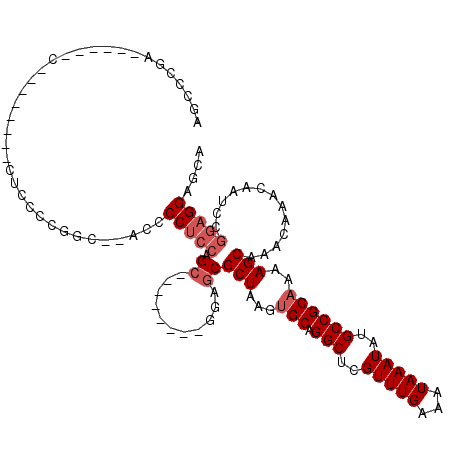
| Consensus MFE | -17.53 |
| Energy contribution | -18.37 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.602441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
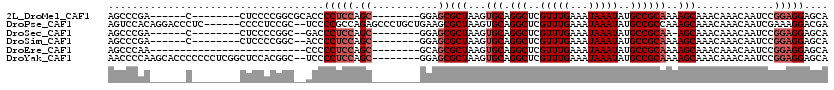
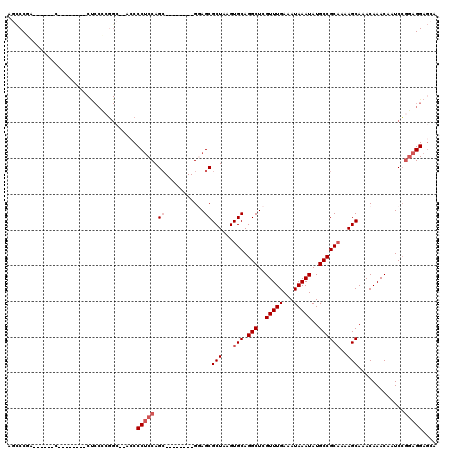
>2L_DroMel_CAF1 12804183 98 - 22407834 AGCCCGA------C--------CUCCCCGGCGCACCCCUCCAGC--------GGAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAAAAGCAAACAAACAAUCCGGAGGAGCA .......------.--------((((((((.((.((.(....).--------)).))(((...(((.(((..(((((...)))))..))))))..)))...........)))).)))).. ( -29.30) >DroPse_CAF1 98617 112 - 1 AGUCCACAGGACCCUC------CCCCUCCGC--UCCCCGCCAGAGCCCUGCUGAAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCCAAAGCAAACAAACAAUCGAAAGGACGA .((((...(((.....------....)))((--((.......))))..((((...((((.....)).(((..(((((...)))))..)))))...))))...............)))).. ( -26.60) >DroSec_CAF1 69863 95 - 1 AGCCCGA------C--------CUCCCCGGC--GACCCUCCAGC--------GGAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAA-AGCAAACAAACAAUCCGGAGGAGCA .......------.--------((((((((.--....((((...--------)))).(((...(((.(((..(((((...)))))..)))))).-)))...........)))).)))).. ( -27.80) >DroSim_CAF1 72039 96 - 1 AGCCCGA------C--------CUCCCCGGC--ACCCCUCCAGC--------GGAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAAAAGCAAACAAACAAUCCGGAGGAGCA .......------.--------((((((((.--....((((...--------)))).(((...(((.(((..(((((...)))))..))))))..)))...........)))).)))).. ( -28.20) >DroEre_CAF1 72203 86 - 1 AGCCCAA--------------------------CCCCCUCCAGC--------GCAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAAAAGCAAACAAACAAUCCGGAGGAGCA .......--------------------------...(((((.((--------...))(((...(((.(((..(((((...)))))..))))))..))).............))))).... ( -22.00) >DroYak_CAF1 73047 110 - 1 AACCCCAAGCACCCCCCCUCGGCUCCACGGC--UCCCCUCCAGC--------GGAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAAAAGCAAACAAACAAUCCGGAGGAGCA .....................(((((.((((--(((.(....).--------)))))(((...(((.(((..(((((...)))))..))))))..)))............))..))))). ( -32.50) >consensus AGCCCGA______C________CUCCCCGGC__ACCCCUCCAGC________GGAGCGCUAAGUGCAGGCUCGUUUGAAAUAAAUAUGCCGCAAAAGCAAACAAACAAUCCGGAGGAGCA ....................................(((((.((...........))(((...(((.(((..(((((...)))))..))))))..))).............))))).... (-17.53 = -18.37 + 0.83)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:15 2006