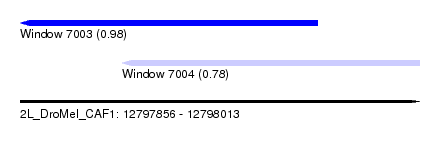
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,797,856 – 12,798,013 |
| Length | 157 |
| Max. P | 0.980291 |

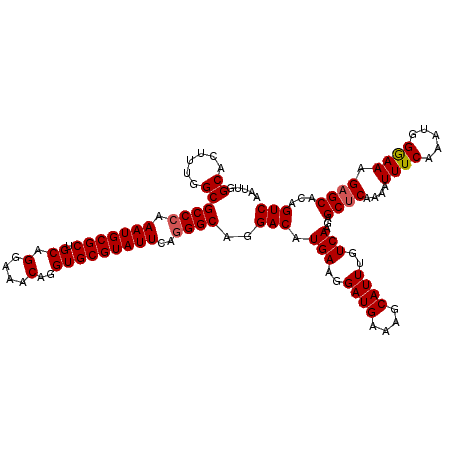
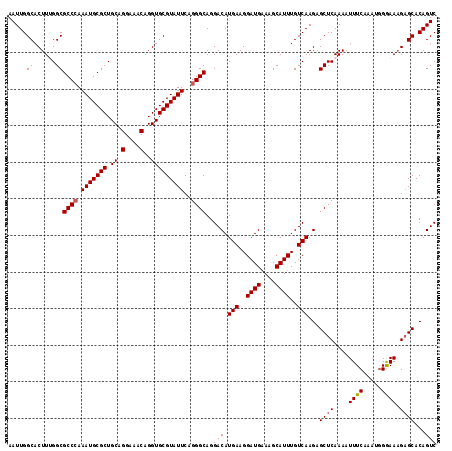
| Location | 12,797,856 – 12,797,973 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 96.58 |
| Mean single sequence MFE | -33.54 |
| Consensus MFE | -30.18 |
| Energy contribution | -30.34 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980291 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12797856 117 - 22407834 AAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUUUGUCAAGAGCUCAAAAUUUCAACUGGGAAUGAGCACAGUC .....((......))((((.(((((((.((.(....)..)))))))))..))))..(((.(((..((((....))))..)))...(((((...(..(....)..).)))))...))) ( -34.00) >DroSec_CAF1 63675 117 - 1 AAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUUUGUCAAGAGCUCAAAAUUUCAAAUGGAAAAGAGCACAGUC .((((((..((((((((((.(((((((.((.(....)..)))))))))..))))......(((..((((....))))..)))...)).)))).((((.....))))...)).)))). ( -32.80) >DroSim_CAF1 65802 117 - 1 AAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUUUGUCAAGAGCUCAAAAUUUCAAAUGGAAAAGAGCACAGUC .((((((..((((((((((.(((((((.((.(....)..)))))))))..))))......(((..((((....))))..)))...)).)))).((((.....))))...)).)))). ( -32.80) >DroEre_CAF1 65632 117 - 1 AAUUGGCACUUAGGCGCCGAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGGAAGCAUUUGUCAAGAGCUCAAAAUUUCUAAUGGGAAAGUGCACAGUC .((((((((((.((((((((((((((((...(....).))))))).)))..)))......(((..((((....))))..)))...))).....(..(....)..))))))).)))). ( -35.70) >DroYak_CAF1 66269 117 - 1 AAUUGGCACUUAGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUUUGUCAAGAGCUCAAAAUUUCAAAUAGGAAAGAGCACAGUC .....((......))((((.(((((((.((.(....)..)))))))))..))))..(((.(((..((((....))))..)))...((((....((((......))))))))...))) ( -32.40) >consensus AAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUUUGUCAAGAGCUCAAAAUUUCAAAUGGGAAAGAGCACAGUC .....((......))((((.(((((((.((.(....)..)))))))))..))))..(((.(((..((((....))))..)))...((((....((((.....)))).))))...))) (-30.18 = -30.34 + 0.16)

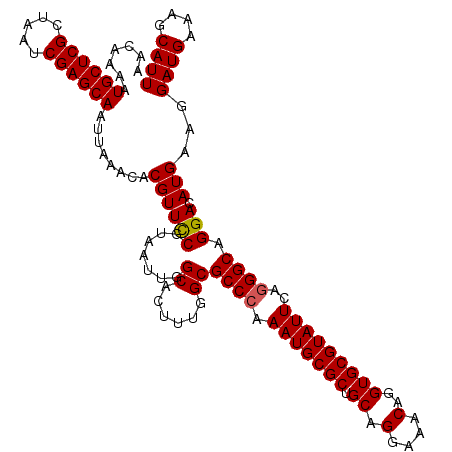
| Location | 12,797,896 – 12,798,013 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 98.29 |
| Mean single sequence MFE | -32.80 |
| Consensus MFE | -30.32 |
| Energy contribution | -30.28 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779478 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12797896 117 - 22407834 AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUCCGUAAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUU ..((((.((((((.....)))))).........((.(((....))).))))))((((((.(((((((.((.(....)..)))))))))..))))....(((.....)))...))... ( -33.20) >DroSec_CAF1 63715 117 - 1 AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUCCGUAAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUU ..((((.((((((.....)))))).........((.(((....))).))))))((((((.(((((((.((.(....)..)))))))))..))))....(((.....)))...))... ( -33.20) >DroSim_CAF1 65842 117 - 1 AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUCCGUAAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUU ..((((.((((((.....)))))).........((.(((....))).))))))((((((.(((((((.((.(....)..)))))))))..))))....(((.....)))...))... ( -33.20) >DroEre_CAF1 65672 117 - 1 AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUUCGUAAUUGGCACUUAGGCGCCGAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGGAAGCAUU .......((((((.....)))))).........(((((.......((((((.(((((......)))))...(....)))))))...((((.(......).)))).....)))))... ( -32.20) >DroYak_CAF1 66309 117 - 1 AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUUCGUAAUUGGCACUUAGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUU .......((((((.....))))))......((..((((((.....((......))((((.(((((((.((.(....)..)))))))))..)))).....))))))..))........ ( -32.20) >consensus AACAAAAUGCUCGCUAAUCGAGCAAUUAAACACGUUCCGUAAUUGGCACUUUGGCGCCCAAAUGCGCUGCAGGAAACAGGUGCGUAUUCAGGGCAGGACAUGAAGGAUGAAAGCAUU .......((((((.....))))))........((((((.......((......))((((.(((((((.((.(....)..)))))))))..)))).))).)))...((((....)))) (-30.32 = -30.28 + -0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:07 2006