
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,763,163 – 12,763,260 |
| Length | 97 |
| Max. P | 0.997212 |

| Location | 12,763,163 – 12,763,260 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 93.18 |
| Mean single sequence MFE | -33.88 |
| Consensus MFE | -26.14 |
| Energy contribution | -27.50 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.878195 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
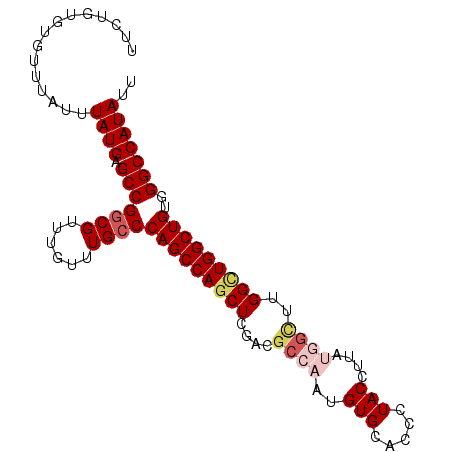
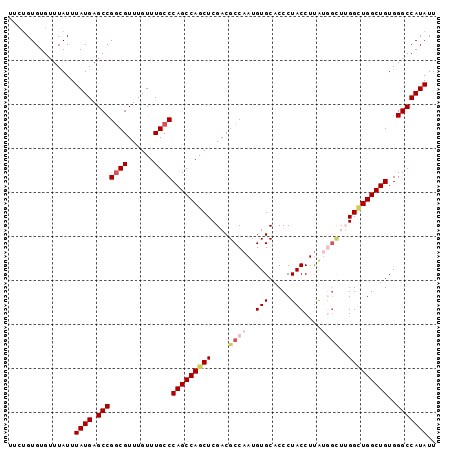
>2L_DroMel_CAF1 12763163 97 + 22407834 UUGUGUGUGUUUAUUUAUGAGCCGGCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUUAUGGCUUGGCUGGCUGUGGGCCAUAUU ...............((((.(((((((......))))((((((((.(((.((((..(((.....)))....)))))))))))))))..))))))).. ( -35.80) >DroSec_CAF1 40317 97 + 1 UUCUGUGUGUUUAUUUAUGAGCCGCCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUUAUGGCUUGGCUGGCUGUGGGCCAUAUU ....(((.(((((....)))))))).........(((((((((((.(((.((((..(((.....)))....)))))))))))))..)))))...... ( -31.60) >DroSim_CAF1 40570 97 + 1 UUCUGUGUGUUUAUUUAUGAGCCGGCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUCAUGGCUUGGCUGGCUGUGGGCCAUAUU ...............((((.(((((((......))))((((((((.(((.((((.((.(........).)))))))))))))))))..))))))).. ( -36.50) >DroEre_CAF1 41144 95 + 1 UUCGGUGUGUUUAUUUAUGAGCCGGCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUUG--GUAGGGUUGGCUGUGGGCCAUAUU ...............((((.(((((((......))))((((((((((...(((((.(((.....))).)))--)).))))))))))..))))))).. ( -36.80) >DroYak_CAF1 41419 92 + 1 UUCUGUGUGUUUAUUUAUGAGCCGGCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUUG--U---GGGUGGCUGUGGGCCAUAUU ...............((((.(((((((((.(((.((...)).)))..))))))......(((((.......--.---)))))......))))))).. ( -28.70) >consensus UUCUGUGUGUUUAUUUAUGAGCCGGCGUUUGUUUGCCCAGCCAGCUCGACGCCAAUGUGCACCCUACCUUAUGGCUUGGCUGGCUGUGGGCCAUAUU ...............((((.(((((((......))))(((((((((....((((..(((.....)))....))))..)))))))))..))))))).. (-26.14 = -27.50 + 1.36)

| Location | 12,763,163 – 12,763,260 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 93.18 |
| Mean single sequence MFE | -33.82 |
| Consensus MFE | -26.98 |
| Energy contribution | -28.26 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.56 |
| Structure conservation index | 0.80 |
| SVM decision value | 2.82 |
| SVM RNA-class probability | 0.997212 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
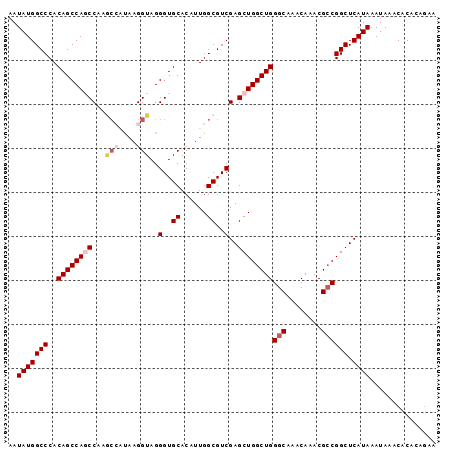
>2L_DroMel_CAF1 12763163 97 - 22407834 AAUAUGGCCCACAGCCAGCCAAGCCAUAAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGCCGGCUCAUAAAUAAACACACACAA ..(((((((..(((((((((..((((....((.....))....))))...).))))))))(((........)))))).))))............... ( -35.70) >DroSec_CAF1 40317 97 - 1 AAUAUGGCCCACAGCCAGCCAAGCCAUAAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGGCGGCUCAUAAAUAAACACACAGAA ..(((((((..(((((((((..((((....((.....))....))))...).)))))))).((.........))))).))))............... ( -31.50) >DroSim_CAF1 40570 97 - 1 AAUAUGGCCCACAGCCAGCCAAGCCAUGAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGCCGGCUCAUAAAUAAACACACAGAA ..(((((((..(((((((((..((((....((.....))....))))...).))))))))(((........)))))).))))............... ( -35.70) >DroEre_CAF1 41144 95 - 1 AAUAUGGCCCACAGCCAACCCUAC--CAAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGCCGGCUCAUAAAUAAACACACCGAA ..(((((((..(((((.(((((((--....))))))).......(((.....))))))))(((........)))))).))))............... ( -36.60) >DroYak_CAF1 41419 92 - 1 AAUAUGGCCCACAGCCACCC---A--CAAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGCCGGCUCAUAAAUAAACACACAGAA ..(((((((..(((((((((---.--.......)))).......(((.....))))))))(((........)))))).))))............... ( -29.60) >consensus AAUAUGGCCCACAGCCAGCCAAGCCAUAAGGUAGGGUGCACAUUGGCGUCGAGCUGGCUGGGCAAACAAACGCCGGCUCAUAAAUAAACACACAGAA ..(((((((..((((((((...(((....)))..(..((......))..)..))))))))(((........)))))).))))............... (-26.98 = -28.26 + 1.28)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:45 2006