
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,606,169 – 12,606,269 |
| Length | 100 |
| Max. P | 0.997689 |

| Location | 12,606,169 – 12,606,269 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 95.50 |
| Mean single sequence MFE | -20.29 |
| Consensus MFE | -18.13 |
| Energy contribution | -18.05 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935721 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
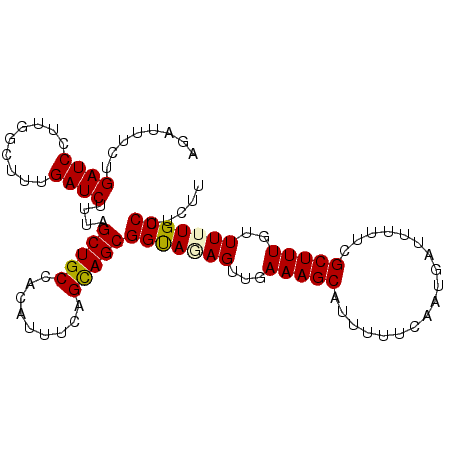
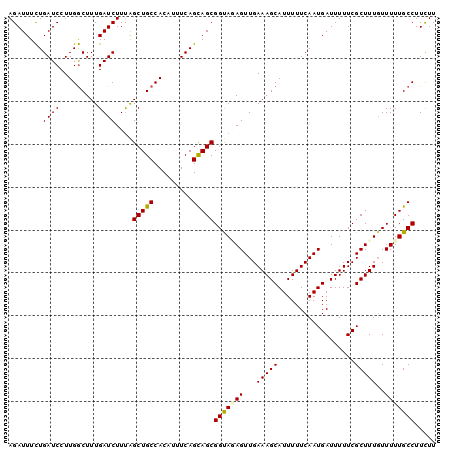
>2L_DroMel_CAF1 12606169 100 + 22407834 AGAUUUCUGAUCCUUGGCUUUGAUCUUUAGCUGCCACAUUUCAGCAGCGGUAUAGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUGUUUUUGCCUUCUU ...............(((...(((.....((((........))))(((((....((((((((...))))))))......)))))..)))...)))..... ( -18.10) >DroSec_CAF1 172996 100 + 1 AGAUUUCUGAUCCUUGGCUUUGAUCUUUAGCUGCCACAUUUCAGCAGCGGUAGAGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUUUUUUUGCCUUCUU ........((((.........))))....(((((.........)))))(((((((..((((((..................)))))).)))))))..... ( -20.07) >DroSim_CAF1 177000 100 + 1 AGAUUUCUGAUCCUUGGCUUUGAUCUUUAGCUGCCACAUUUCAGUAGCGGUAGAGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUGUUUUUGCCUUCUU (((((..(((......(((((.(.(((((.((((.((......)).))))))))).).))))).....)))..)))))...((.........))...... ( -19.20) >DroEre_CAF1 171219 100 + 1 AGAUUUCUGAUCCUUGGCUUUGAUCUUCAGCUGCCACAUUUCAGCAGCGGCAAAGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUGUUUUUGCCUUUUU .......(((....((((.(((.....)))..))))....)))((((..(((((((.(((((((((....)))).))))).)))))))..))))...... ( -24.50) >DroAna_CAF1 171930 99 + 1 AGAUUUUUGAUCCUUGACUUUGAUCUUUGGCUGCCACAUUUCAGCAGCGGCA-AGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUGUUUUUGCCUUUUU ........((((.........))))....(((((.........)))))((((-((...(((((..................)))))...))))))..... ( -19.57) >consensus AGAUUUCUGAUCCUUGGCUUUGAUCUUUAGCUGCCACAUUUCAGCAGCGGUAGAGUUGAAAGCAUUUUUCAAUGAUUUUUCGCUUUGUUUUUGCCUUCUU ........((((.........))))....(((((.........)))))(((((((...(((((..................)))))..)))))))..... (-18.13 = -18.05 + -0.08)

| Location | 12,606,169 – 12,606,269 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 95.50 |
| Mean single sequence MFE | -20.98 |
| Consensus MFE | -17.74 |
| Energy contribution | -17.78 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.91 |
| SVM RNA-class probability | 0.997689 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
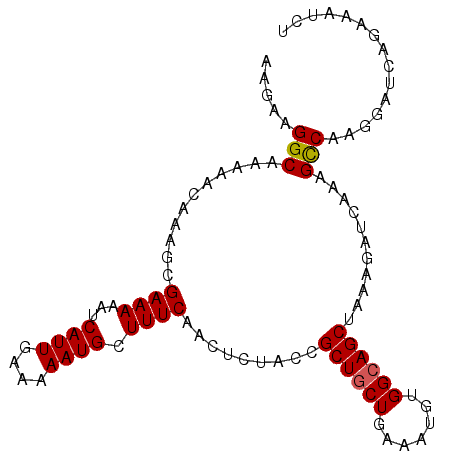
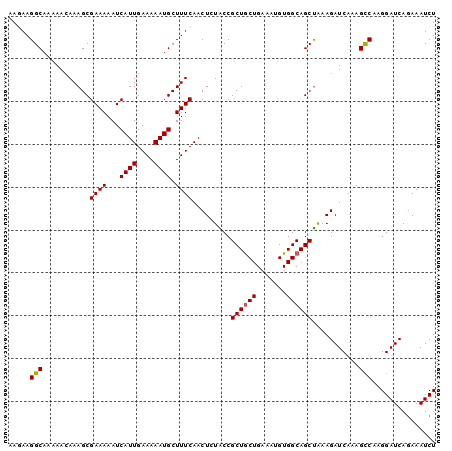
>2L_DroMel_CAF1 12606169 100 - 22407834 AAGAAGGCAAAAACAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACUAUACCGCUGCUGAAAUGUGGCAGCUAAAGAUCAAAGCCAAGGAUCAGAAAUCU .....(((...........((((...((((....)))).)))).........((((((.......))))))...........)))............... ( -19.50) >DroSec_CAF1 172996 100 - 1 AAGAAGGCAAAAAAAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACUCUACCGCUGCUGAAAUGUGGCAGCUAAAGAUCAAAGCCAAGGAUCAGAAAUCU ..((((((...........((....))(((....)))))))))...(((...((((((.......))))))....((((.........)))))))..... ( -20.30) >DroSim_CAF1 177000 100 - 1 AAGAAGGCAAAAACAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACUCUACCGCUACUGAAAUGUGGCAGCUAAAGAUCAAAGCCAAGGAUCAGAAAUCU ..((((((...........((....))(((....)))))))))...(((...(((((......))))).......((((.........)))))))..... ( -18.20) >DroEre_CAF1 171219 100 - 1 AAAAAGGCAAAAACAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACUUUGCCGCUGCUGAAAUGUGGCAGCUGAAGAUCAAAGCCAAGGAUCAGAAAUCU .....((((((...(((((((....))(((....))))))))....))))))((((((.......))))))....((((.........))))........ ( -23.90) >DroAna_CAF1 171930 99 - 1 AAAAAGGCAAAAACAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACU-UGCCGCUGCUGAAAUGUGGCAGCCAAAGAUCAAAGUCAAGGAUCAAAAAUCU .....(((((....(((((((....))(((....))))))))....)-))))((((((.......))))))....((((.........))))........ ( -23.00) >consensus AAGAAGGCAAAAACAAAGCGAAAAAUCAUUGAAAAAUGCUUUCAACUCUACCGCUGCUGAAAUGUGGCAGCUAAAGAUCAAAGCCAAGGAUCAGAAAUCU .....(((...........((((...((((....)))).)))).........((((((.......))))))...........)))............... (-17.74 = -17.78 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:16:18 2006