
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,553,691 – 12,553,801 |
| Length | 110 |
| Max. P | 0.996438 |

| Location | 12,553,691 – 12,553,801 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.63 |
| Mean single sequence MFE | -30.77 |
| Consensus MFE | -10.69 |
| Energy contribution | -11.80 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.35 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585115 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
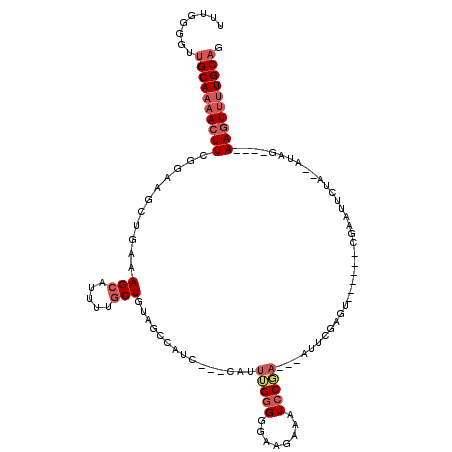
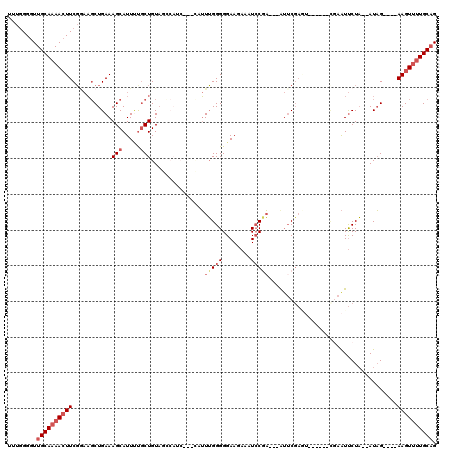
>2L_DroMel_CAF1 12553691 110 + 22407834 UUUGGGGUUGCAAAACUUCGGAAGCUGAAAGCAUUUUGCUGUAGCCAUCGCAU-UUUGGGGGAAGAAAUCCGA---AUUCGAGUCCAAACCGAAUUCUA--AUAG----AAGUUUUGCAG ........((((((((((((((.((((..(((.....))).))))..((.(..-.....).)).....)))((---(((((.((....)))))))))..--...)----)))))))))). ( -33.50) >DroSec_CAF1 125847 109 + 1 UUUGGGGUUGCAAAACUUCGGAAGCUGAAAGCAUUGUGCUGUAGCCAUC--GCCUUUGGGGGAAGAAAUCCGA---GUUCGAGUCCAAACCAAAUUCUA--AUAG----AAGUUUUGCAG ........(((((((((((((..((((..(((.....))).))))....--.))(((((.(((.(((......---.)))...)))...))))).....--...)----)))))))))). ( -32.00) >DroSim_CAF1 128622 111 + 1 UUUGGGGUUGCAAAACUUCGGAAGCUGAAAGCAUUUUGCUGUAGCCAUCUCACAUUUGGGGGAAGAAAUCCGA---GUUCGAAUACGAACCGAAUUCUA--AUAG----AAGUUUUGCAG ........((((((((((((((.((((..(((.....))).))))..(((((....))))).......)))((---(((((.........)))))))..--...)----)))))))))). ( -34.50) >DroEre_CAF1 124780 103 + 1 UUUGGGGUUGCAAAACUUCGGAAGCUGAAAGCAUUUUGCUGUACGCAGC--GCAUUUGGGGGAAGAAAUGCGAGAAAUGCGA---------GAAUUCUA--AUAG----AAGUUUUGCAG ........(((((((((((((((.......(((((((((((....))))--((((((........))))))..)))))))..---------...)))).--...)----)))))))))). ( -34.50) >DroYak_CAF1 129908 96 + 1 UUUGGGGUUGCAAAACUUCGCAAGCUGAAAGCAUUUCGCUGU-----------AUUUGGGGGAAGAAAUCCUA---AGUCGAGU------UGAAUUCUAGAAUAG----AAGUUUUGCAG ........(((((((((((.....(((((..((.((((....-----------((((((((.......)))))---))))))).------))..))).))....)----)))))))))). ( -28.20) >DroAna_CAF1 125994 94 + 1 UUUAGGGUUGCAAAAGUUCCACUCCUGAAAGCAUUCUCCUGUACG---------AUGGGGGGAA---GUCCUU---AUUCC---------UUAGUCCUC--AUUGAAGCAAGUCCUGCUG ((((((((.((....))...)).))))))((((....(.(((.((---------((((((((((---......---.))))---------.....))))--))))..))).)...)))). ( -21.90) >consensus UUUGGGGUUGCAAAACUUCGGAAGCUGAAAGCAUUUUGCUGUAGCCAUC___CAUUUGGGGGAAGAAAUCCGA___AUUCGAGU______CGAAUUCUA__AUAG____AAGUUUUGCAG ........((((((((((...........(((.....)))...............(((((........)))))....................................)))))))))). (-10.69 = -11.80 + 1.11)

| Location | 12,553,691 – 12,553,801 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.63 |
| Mean single sequence MFE | -27.33 |
| Consensus MFE | -11.44 |
| Energy contribution | -11.83 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.42 |
| SVM decision value | 2.70 |
| SVM RNA-class probability | 0.996438 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
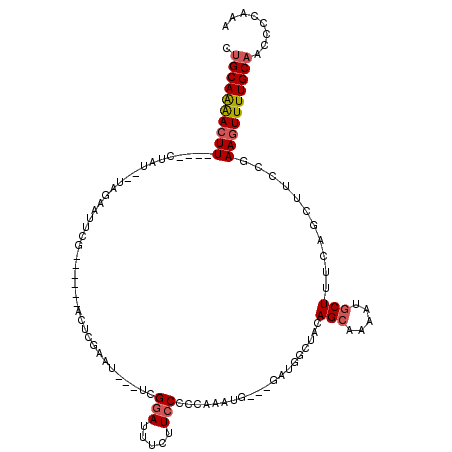
>2L_DroMel_CAF1 12553691 110 - 22407834 CUGCAAAACUU----CUAU--UAGAAUUCGGUUUGGACUCGAAU---UCGGAUUUCUUCCCCCAAA-AUGCGAUGGCUACAGCAAAAUGCUUUCAGCUUCCGAAGUUUUGCAACCCCAAA .((((((((((----(...--..(((((((((....)).)))))---))(((.....)))......-....(..((((..(((.....)))...))))..))))))))))))........ ( -32.30) >DroSec_CAF1 125847 109 - 1 CUGCAAAACUU----CUAU--UAGAAUUUGGUUUGGACUCGAAC---UCGGAUUUCUUCCCCCAAAGGC--GAUGGCUACAGCACAAUGCUUUCAGCUUCCGAAGUUUUGCAACCCCAAA .((((((((((----(...--..((((((((((((....)))))---.))))))).........(((((--(.((((....)).)).))))))........)))))))))))........ ( -30.20) >DroSim_CAF1 128622 111 - 1 CUGCAAAACUU----CUAU--UAGAAUUCGGUUCGUAUUCGAAC---UCGGAUUUCUUCCCCCAAAUGUGAGAUGGCUACAGCAAAAUGCUUUCAGCUUCCGAAGUUUUGCAACCCCAAA .((((((((((----(...--..((((((((((((....)))))---.)))))))................(..((((..(((.....)))...))))..))))))))))))........ ( -33.40) >DroEre_CAF1 124780 103 - 1 CUGCAAAACUU----CUAU--UAGAAUUC---------UCGCAUUUCUCGCAUUUCUUCCCCCAAAUGC--GCUGCGUACAGCAAAAUGCUUUCAGCUUCCGAAGUUUUGCAACCCCAAA .((((((((((----(...--..((....---------))((((((...((((((........))))))--((((....)))).))))))...........)))))))))))........ ( -29.60) >DroYak_CAF1 129908 96 - 1 CUGCAAAACUU----CUAUUCUAGAAUUCA------ACUCGACU---UAGGAUUUCUUCCCCCAAAU-----------ACAGCGAAAUGCUUUCAGCUUGCGAAGUUUUGCAACCCCAAA .((((((((((----(..............------........---..(((.....))).......-----------...((((...((.....)))))))))))))))))........ ( -21.20) >DroAna_CAF1 125994 94 - 1 CAGCAGGACUUGCUUCAAU--GAGGACUAA---------GGAAU---AAGGAC---UUCCCCCCAU---------CGUACAGGAGAAUGCUUUCAGGAGUGGAACUUUUGCAACCCUAAA ..((((((.(..((((.((--((((.....---------((((.---......---))))..)).)---------)))....((((....)))).))))..)...))))))......... ( -17.30) >consensus CUGCAAAACUU____CUAU__UAGAAUUCG______ACUCGAAU___UCGGAUUUCUUCCCCCAAAUG___GAUGGCUACAGCAAAAUGCUUUCAGCUUCCGAAGUUUUGCAACCCCAAA .((((((((((......................................(((.....)))....................(((.....)))...........))))))))))........ (-11.44 = -11.83 + 0.39)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:16:05 2006