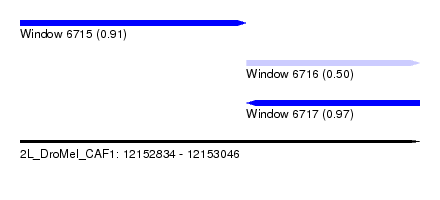
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,152,834 – 12,153,046 |
| Length | 212 |
| Max. P | 0.966014 |

| Location | 12,152,834 – 12,152,954 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.83 |
| Mean single sequence MFE | -23.63 |
| Consensus MFE | -19.57 |
| Energy contribution | -19.89 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911291 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12152834 120 + 22407834 AAAUAAAUACUGAAAGCAUAAAUAAAAUAGGUGGGAGUGGGUGGAAAACACUUGCAGAACCGACCCAUUUCCACGUCAGCACAGUGAAAGCGAGCGAAAUGUAAAUAUCAAAUAAAACGC ...............((((...........((((((((((((((...............)).))))))))))))....((...((....))..))...)))).................. ( -26.66) >DroSec_CAF1 38055 114 + 1 ------UUACUGAAAGCAUAAAUAAAAUAGGUGGGAAUGGGUGGAAAACACUUGCAGAACCGACCCAUUUUCUCGUCAGCACAGUGAAAGCGAGCAAAAUGUAAAUAUCAAAUAAAACGC ------.........((............(((.((..((((((.....))))..))...)).)))(((((((((((...(.....)...))))).)))))).................)) ( -21.30) >DroSim_CAF1 34705 120 + 1 AAAUAAAUACUGAAAGCAUAAAUAAAAUAGGUGGGAAUGGGUGGAAAGCACUUGCAGAACCGACCCAUUUCCACGUCAGCACAGUGAAAGCGAGCGAAAUGUAAAUAUCAAAUAAAACGC ...............((((...........((((((((((((((...((....))....)).))))))))))))....((...((....))..))...)))).................. ( -29.40) >DroEre_CAF1 33908 111 + 1 AAAUAAAUACUGAAAGCAUAAAUAAAACAAGAGUGAAUGGGUGGAAAACACUUGCAGAAGCGACCCAUUUCCACGUCAGCGCAGUGAAAGCGAGCGAAAUGUAAAUAUCAA--------- .......(((.....((...............(((..((((((((((....((((....))))....))))))).))).))).((....))..)).....)))........--------- ( -21.00) >DroYak_CAF1 28568 120 + 1 AAAUAAAUACUGAAGGCAUAAAUAAAAUAAGAGGGUAUGGGUGGAAAACACUUGCAGAAGCGACCCAUUUCCACGUCAGCACAGUGAAAGCGAGCGAAAUGUAAAUAUCAAAUAAAACGC ..........(((..((((.............(((.(((((((.....)))((((....)))).)))).))).(((..((.........))..)))..)))).....))).......... ( -19.80) >consensus AAAUAAAUACUGAAAGCAUAAAUAAAAUAGGUGGGAAUGGGUGGAAAACACUUGCAGAACCGACCCAUUUCCACGUCAGCACAGUGAAAGCGAGCGAAAUGUAAAUAUCAAAUAAAACGC ...............((((.............((((((((((.(................).)))))))))).(((..((.........))..)))..)))).................. (-19.57 = -19.89 + 0.32)

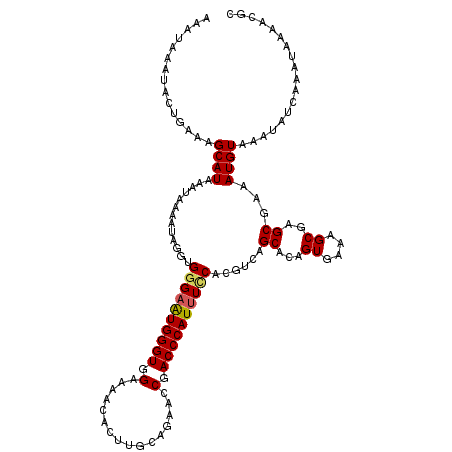
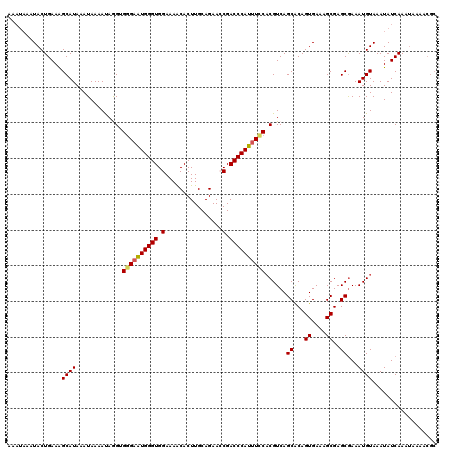
| Location | 12,152,954 – 12,153,046 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 84.13 |
| Mean single sequence MFE | -21.44 |
| Consensus MFE | -15.80 |
| Energy contribution | -16.60 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12152954 92 + 22407834 GUAGCAAAAA--AAAAAGGGAAAUGCUACACAUUUGCGUUUGAGCUGAACUGAAAGGCAUGGAAAGUAAGUUUUGGACUUGAUUUCCAGACCAC ((((((....--...........)))))).....(((.(((.((.....)).))).)))((((((.((((((...)))))).))))))...... ( -22.56) >DroSec_CAF1 38169 94 + 1 GUAGCAAAAAGAAAGAAGGGAAAUGCUACACAUUUGCGUUUGAGCUGAACUGAAAGGCAUGGAAAGUAAGUUUUGGACUUGAUUUCCAGACCAC ((((((.................)))))).....(((.(((.((.....)).))).)))((((((.((((((...)))))).))))))...... ( -22.43) >DroSim_CAF1 34825 94 + 1 GUAGCAAAGAAGAAAAAGGGAAAUGCUACACAUUUGCGUUUGAGCUGAACUGAAAGGCAUGGAAAGUAAGUUUUGGACUUGAUUUCCAGACCAC ((((((.................)))))).....(((.(((.((.....)).))).)))((((((.((((((...)))))).))))))...... ( -22.43) >DroEre_CAF1 34019 71 + 1 ----------------------AUGCUACACAUUUGCGUUUGAGCUGAGCUGAAAGGCAUGGA-AGUAAGUUUUGGACUUGAUUUCCACGCCAC ----------------------.(((.........)))(((.(((...))).)))(((.((((-((((((((...))))).))))))).))).. ( -19.30) >DroYak_CAF1 28688 84 + 1 GUAGGGA---------AAGGAAAUGCCACACAUUUGCGUUUGAGCUGAGCUGAAAGGCAUGGA-AGUAAGUUUUGGACUUGAUUUCCACACCAC ...((..---------..((.....)).......(((.(((.(((...))).))).)))((((-((((((((...))))).)))))))..)).. ( -20.50) >consensus GUAGCAAA_A__AA_AAGGGAAAUGCUACACAUUUGCGUUUGAGCUGAACUGAAAGGCAUGGAAAGUAAGUUUUGGACUUGAUUUCCAGACCAC ..................((..............(((.(((.((.....)).))).)))((((((.((((((...)))))).))))))..)).. (-15.80 = -16.60 + 0.80)

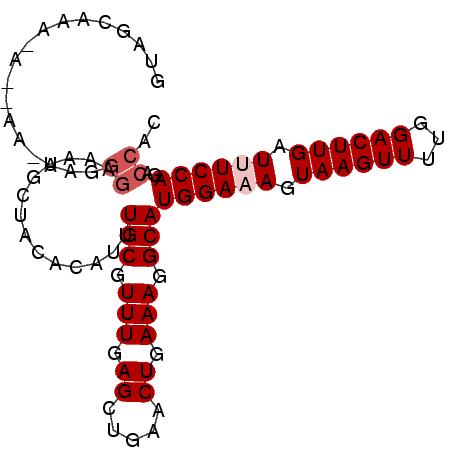
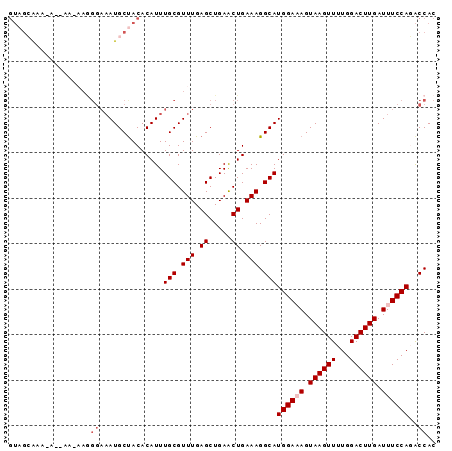
| Location | 12,152,954 – 12,153,046 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 84.13 |
| Mean single sequence MFE | -19.08 |
| Consensus MFE | -17.20 |
| Energy contribution | -17.28 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.966014 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12152954 92 - 22407834 GUGGUCUGGAAAUCAAGUCCAAAACUUACUUUCCAUGCCUUUCAGUUCAGCUCAAACGCAAAUGUGUAGCAUUUCCCUUUUU--UUUUUGCUAC ..(((.((((((..((((.....))))..)))))).)))..........((......))......((((((...........--....)))))) ( -18.76) >DroSec_CAF1 38169 94 - 1 GUGGUCUGGAAAUCAAGUCCAAAACUUACUUUCCAUGCCUUUCAGUUCAGCUCAAACGCAAAUGUGUAGCAUUUCCCUUCUUUCUUUUUGCUAC ..(((.((((((..((((.....))))..)))))).)))..........((......))......((((((.................)))))) ( -18.63) >DroSim_CAF1 34825 94 - 1 GUGGUCUGGAAAUCAAGUCCAAAACUUACUUUCCAUGCCUUUCAGUUCAGCUCAAACGCAAAUGUGUAGCAUUUCCCUUUUUCUUCUUUGCUAC ..(((.((((((..((((.....))))..)))))).)))..........((......))......((((((.................)))))) ( -18.63) >DroEre_CAF1 34019 71 - 1 GUGGCGUGGAAAUCAAGUCCAAAACUUACU-UCCAUGCCUUUCAGCUCAGCUCAAACGCAAAUGUGUAGCAU---------------------- ..(((((((((...((((.....))))..)-))))))))..........(((...((((....)))))))..---------------------- ( -21.00) >DroYak_CAF1 28688 84 - 1 GUGGUGUGGAAAUCAAGUCCAAAACUUACU-UCCAUGCCUUUCAGCUCAGCUCAAACGCAAAUGUGUGGCAUUUCCUU---------UCCCUAC ..(((((((((...((((.....))))..)-))))))))..........(((...((((....)))))))........---------....... ( -18.40) >consensus GUGGUCUGGAAAUCAAGUCCAAAACUUACUUUCCAUGCCUUUCAGUUCAGCUCAAACGCAAAUGUGUAGCAUUUCCCUU_UU__U_UUUGCUAC ..(((.((((((..((((.....))))..)))))).)))..........(((...((((....)))))))........................ (-17.20 = -17.28 + 0.08)

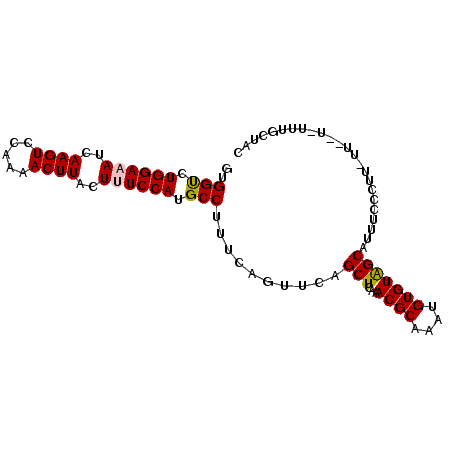
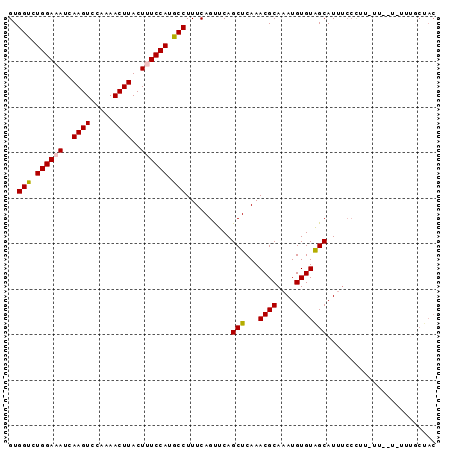
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:13:33 2006