| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,116,462 – 12,116,560 |
| Length | 98 |
| Max. P | 0.737467 |

| Location | 12,116,462 – 12,116,560 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.30 |
| Mean single sequence MFE | -30.18 |
| Consensus MFE | -18.34 |
| Energy contribution | -19.18 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.737467 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12116462 98 + 22407834 ACUGCUUCUUCUUGCCACUUCUUGGAGCUUGACGUGACGACAUGCAAUCGCAAAGAAGGUA--------GAAAGAAGA------A----GCU----AUAAGAAGCGGUAGAACAACACCA ...((((((((((....((((((...(.(((.((((....))))))).)...))))))...--------..)))))))------)----)).----.........(((........))). ( -29.00) >DroSec_CAF1 18580 91 + 1 UCUGCUUCUUCUUGCCACUUCUUGGAGCUUGACGUGACGACAUGCAAUCGCAAAGAAGGUA--------GAAAGAAGAAG---------------------AAGCGGUAGAACAACACCA .((((((((((((.(..((((((...(.(((.((((....))))))).)...))))))...--------)....))))))---------------------))))))............. ( -27.10) >DroSim_CAF1 18413 101 + 1 UCUGCUUCUUCUUGCCACUUCUUGGAGCUUGACGUGACGACAUGCAAUCGCAAAGAAGGUA--------GAAAGAAGAAGA---A----GCU----AUAAGAAGCGGUAGAACAACACCA ....(((((((((....((((((...(.(((.((((....))))))).)...))))))...--------..))))))))).---.----(((----......)))(((........))). ( -28.20) >DroEre_CAF1 18849 116 + 1 UCUGCUUCUUCUUGCCACUUCUUAGAGCUUGACGUGACGACAUGCAAUCGCAAAGAAGGUAAGAAGGUAGGAAGAAGAAGAUGAAGCGGGCU----AUAACAAGCGGUAGAACAACACCA (((.((((((((((((.((((((.(.(.(((.((((....))))))).).).)))))).......)))))))))))).)))........(((----......)))(((........))). ( -33.40) >DroYak_CAF1 18763 115 + 1 UCUGCUUCUUCUUGCCACUUCUUAGAGCUUCACGUGACGACAUGCAAUCGCAAAGAAGGUA-----GUUGGAAGAAGAAGAUGAAGCUGGCUACAUAUAAGAAGCGGCAGAACAACACCA ((((((.((((((((((((((......((((......((((.(((..((.....))..)))-----))))......))))..)))).)))).......)))))).))))))......... ( -33.21) >consensus UCUGCUUCUUCUUGCCACUUCUUGGAGCUUGACGUGACGACAUGCAAUCGCAAAGAAGGUA________GAAAGAAGAAGA___A____GCU____AUAAGAAGCGGUAGAACAACACCA (((((((((((((.(..((((((...(.(((.((((....))))))).)...))))))...........).)))))))...........(((..........)))))))))......... (-18.34 = -19.18 + 0.84)


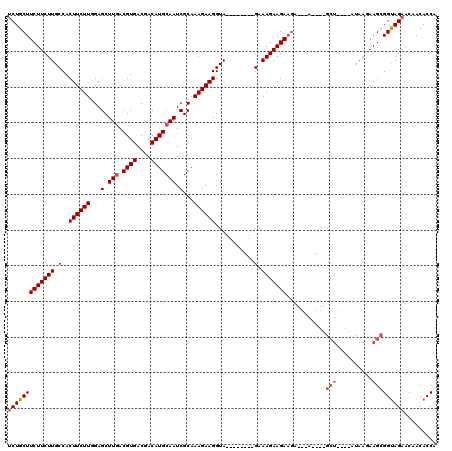
| Location | 12,116,462 – 12,116,560 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.30 |
| Mean single sequence MFE | -28.69 |
| Consensus MFE | -18.46 |
| Energy contribution | -18.90 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.561163 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12116462 98 - 22407834 UGGUGUUGUUCUACCGCUUCUUAU----AGC----U------UCUUCUUUC--------UACCUUCUUUGCGAUUGCAUGUCGUCACGUCAAGCUCCAAGAAGUGGCAAGAAGAAGCAGU .((((......)))).........----.((----(------(((((((.(--------(((.(((((.(.(.(((.((((....))))))).).).))))))))).))))))))))... ( -29.90) >DroSec_CAF1 18580 91 - 1 UGGUGUUGUUCUACCGCUU---------------------CUUCUUCUUUC--------UACCUUCUUUGCGAUUGCAUGUCGUCACGUCAAGCUCCAAGAAGUGGCAAGAAGAAGCAGA .((((......))))((((---------------------((((((....(--------(((.(((((.(.(.(((.((((....))))))).).).))))))))).))))))))))... ( -27.10) >DroSim_CAF1 18413 101 - 1 UGGUGUUGUUCUACCGCUUCUUAU----AGC----U---UCUUCUUCUUUC--------UACCUUCUUUGCGAUUGCAUGUCGUCACGUCAAGCUCCAAGAAGUGGCAAGAAGAAGCAGA .((((......))))(((......----)))----.---.(((((((((.(--------(((.(((((.(.(.(((.((((....))))))).).).))))))))).))))))))).... ( -27.50) >DroEre_CAF1 18849 116 - 1 UGGUGUUGUUCUACCGCUUGUUAU----AGCCCGCUUCAUCUUCUUCUUCCUACCUUCUUACCUUCUUUGCGAUUGCAUGUCGUCACGUCAAGCUCUAAGAAGUGGCAAGAAGAAGCAGA .((((......))))(((......----))).........(((((((((.((((.((((((.(((...((((((.....)))).))....)))...)))))))))).))))))))).... ( -28.10) >DroYak_CAF1 18763 115 - 1 UGGUGUUGUUCUGCCGCUUCUUAUAUGUAGCCAGCUUCAUCUUCUUCUUCCAAC-----UACCUUCUUUGCGAUUGCAUGUCGUCACGUGAAGCUCUAAGAAGUGGCAAGAAGAAGCAGA ...((((.(((((((((((((((.........((((((((..............-----..........(((((.....)))))...)))))))).))))))))))).))))..)))).. ( -30.83) >consensus UGGUGUUGUUCUACCGCUUCUUAU____AGC____U___UCUUCUUCUUUC________UACCUUCUUUGCGAUUGCAUGUCGUCACGUCAAGCUCCAAGAAGUGGCAAGAAGAAGCAGA .((((......)))).........................(((((((((...................(((....)))....(((((.((.........)).)))))))))))))).... (-18.46 = -18.90 + 0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:13:09 2006