
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,112,552 – 12,112,711 |
| Length | 159 |
| Max. P | 0.999949 |

| Location | 12,112,552 – 12,112,672 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.50 |
| Mean single sequence MFE | -31.06 |
| Consensus MFE | -29.08 |
| Energy contribution | -29.48 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.94 |
| SVM decision value | 4.30 |
| SVM RNA-class probability | 0.999865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12112552 120 - 22407834 UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCGAGCAAUUAGUUUCCCAAUAAGAACAUUCCGAUCAAUAUGAUU (((((((((((((((.(((..((....))...))).)))))))))..(((((((((..(((.((..........)))))))))))))))))))).............((((.....)))) ( -32.50) >DroSec_CAF1 14605 120 - 1 UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCAAGCAAUUAGUUUCCCAAUAAGAACUUUCCGAUCAACAUGAUU (((((((((((((((.(((..((....))...))).)))))))))..(((((((((....(((..........)))...))))))))))))))).............((((.....)))) ( -32.00) >DroSim_CAF1 14437 120 - 1 UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCAAGCAAUUAGUUUCCCAAUAAGAACUUUCCGAUCAAUAUGAUU (((((((((((((((.(((..((....))...))).)))))))))..(((((((((....(((..........)))...))))))))))))))).............((((.....)))) ( -32.00) >DroEre_CAF1 14664 120 - 1 UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUGCUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACUGACCAAUUAGUUUCCCAAUGAGACCUUUCCCAUCAAUAUGAUU (((((((((((((((.(((..((....))...))).)))))))))((((((((((.(.(((((..........))).))))))))))))).............))))))........... ( -29.80) >DroYak_CAF1 14801 120 - 1 UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUGCUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACUGACCAAUUAGUUUCCCAAUGAGACCUUUCCCUUCAAUAUGAUU ((((((((((((..((((((......)))((((((((((((((......))).)))))......)))))).......)))..)))))))))))).......................... ( -29.00) >consensus UGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCGAGCAAUUAGUUUCCCAAUAAGAACUUUCCGAUCAAUAUGAUU (((((((((((((((.(((..((....))...))).)))))))))..(((((((((....(((..........)))...))))))))))))))).......................... (-29.08 = -29.48 + 0.40)


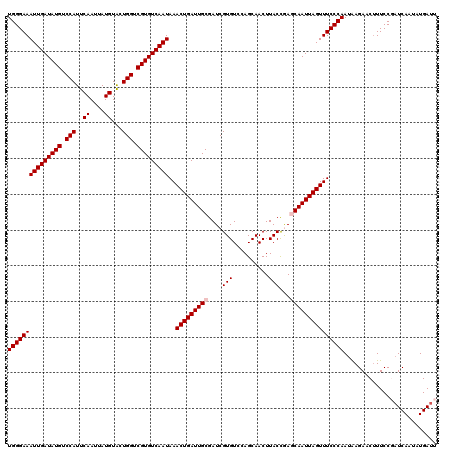
| Location | 12,112,592 – 12,112,711 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.82 |
| Mean single sequence MFE | -37.92 |
| Consensus MFE | -36.16 |
| Energy contribution | -35.92 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.95 |
| SVM decision value | 4.78 |
| SVM RNA-class probability | 0.999949 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12112592 119 - 22407834 -UUUGGAGGCACUUAAUAAGCUUUGCUGGUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCGAG -.((((((((.........))))(((((((..(((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..)))))..).)))))).....)))). ( -40.60) >DroSec_CAF1 14645 120 - 1 UUUUGGAGGCACUUAAUAAGCUUUGCUGGUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCAAG ..((((((((.........))))(((((((..(((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..)))))..).)))))).....)))). ( -40.90) >DroSim_CAF1 14477 119 - 1 -UUUGGAGGCACUUAAUAAGCUUUGCUGGUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCAAG -.((((((((.........))))(((((((..(((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..)))))..).)))))).....)))). ( -40.90) >DroEre_CAF1 14704 119 - 1 -UUUGGAGGCACUUAAUAAGCUUUUCUGGUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUGCUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACUGAC -.(((((.((.........(((.....)))..(((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..))))))).)))))............ ( -33.90) >DroYak_CAF1 14841 119 - 1 -UUUGGAGGCACUUAAUAAGCUUUUCUGAUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUGCUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACUGAC -...((((((.........))))))..((..((((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..))))))..))(((.......))).. ( -33.30) >consensus _UUUGGAGGCACUUAAUAAGCUUUGCUGGUGUGAUUGUGAUGGGAAAUUGAUAUGUCCAUUCAAUUAUGUACUGGUCGUGUCAAUAAACUGAUUGCGAUCGUGUCCAGCAACUUACCGAG ..((((((((.........))))((((((..((((((..((.((..(((((((((.(((..((....))...))).)))))))))...)).))..))))))..).))))).....)))). (-36.16 = -35.92 + -0.24)

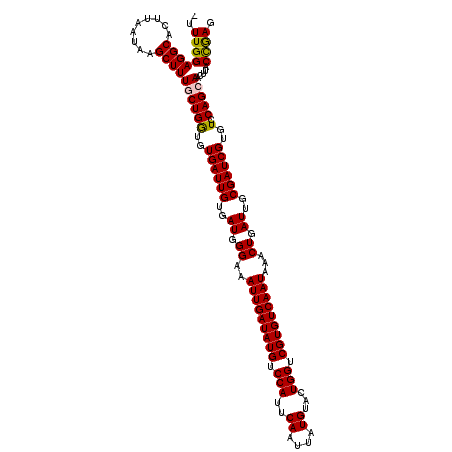

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:58 2006