
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 12,010,373 – 12,010,467 |
| Length | 94 |
| Max. P | 0.886317 |

| Location | 12,010,373 – 12,010,467 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.50 |
| Mean single sequence MFE | -37.48 |
| Consensus MFE | -23.90 |
| Energy contribution | -23.86 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12010373 94 + 22407834 UCCGGAUACCUCCGGAGCGAGGAGCAUGGCAACAAUGUUGCAGUUGCUCGUA-------------------CUCGUACUCCAAGGGGAUGGGAAG----UAGAUGCGUGUGU---GCACA ((((((....))))))(((..(((((..(((((...)))))...)))))...-------------------(((((.(((....))))))))...----....)))(((...---.))). ( -35.30) >DroSec_CAF1 1867 94 + 1 UCCGGAUACCUCCGGAGCGAGGAGCAUGGCAACAAUGUUGCAGUUGCUCAUA-------------------CUCGUACUCCAAGGGGAUGGGAAG----UUGAUGCGUGUGU---GCACA ((((((....))))))(((..(((((..(((((...)))))...)))))...-------------------(((((.(((....))))))))...----....)))(((...---.))). ( -35.30) >DroSim_CAF1 1868 94 + 1 UCCGGAUACCUCCGGAGCGAGGAGCAUGGCAACAAUGUUGCAGUUGCUCGUA-------------------CUCGUACUCCAUGGGGAUGGGAAG----UUGAUGCGUGUGU---GCACA ((((((....))))))(((..(((((..(((((...)))))...)))))...-------------------(((((.(((....))))))))...----....)))(((...---.))). ( -35.30) >DroEre_CAF1 2210 116 + 1 ACCGGAUACCUCCGGAGCGAGGAGCAUGGCAACAAUGUUGCAGUUGUUCGUGCUGGUGCUGGGUGCUUGUGCUCGCACUC-CAGUGCAAUGGAAGCAUGUGGAUGCGUGUGU---GCACA .(((((....)))))......(((((..(((((...)))))...)))))((((..(..((((((((........)))).)-)))..).......((((((....))))))..---)))). ( -47.60) >DroYak_CAF1 2239 91 + 1 ACCGGAUACCUCCGGAGCGAGCAGCAUGGCAACAAUGUUGCAGUUGUUCGUGCU------------------------CC-AAGGGGAAUGGAAG----UAGAUACGUGUGUGCUGCACA .(((....(((..((((((((((((...(((((...))))).)))))))..)))------------------------))-..)))...)))..(----(((.(((....)))))))... ( -33.90) >consensus UCCGGAUACCUCCGGAGCGAGGAGCAUGGCAACAAUGUUGCAGUUGCUCGUA___________________CUCGUACUCCAAGGGGAUGGGAAG____UAGAUGCGUGUGU___GCACA .(((((....))))).(((((.(((...(((((...))))).))).))))).......................................................((((.....)))). (-23.90 = -23.86 + -0.04)

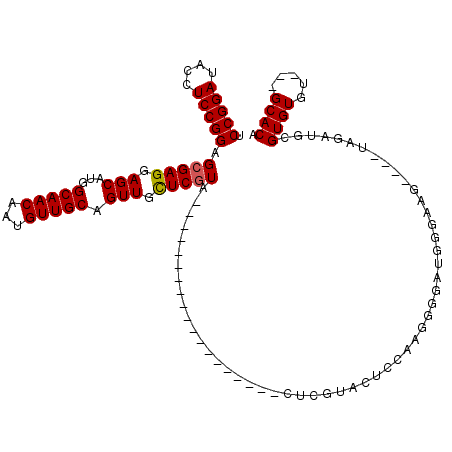
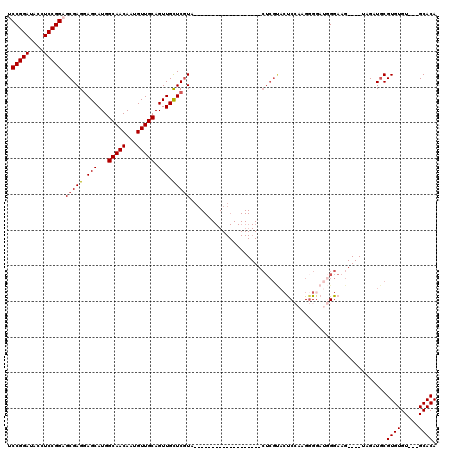
| Location | 12,010,373 – 12,010,467 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.50 |
| Mean single sequence MFE | -30.28 |
| Consensus MFE | -18.70 |
| Energy contribution | -19.50 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.793132 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 12010373 94 - 22407834 UGUGC---ACACACGCAUCUA----CUUCCCAUCCCCUUGGAGUACGAG-------------------UACGAGCAACUGCAACAUUGUUGCCAUGCUCCUCGCUCCGGAGGUAUCCGGA .((((---......)))).((----((..(((......)))))))((((-------------------...(((((...(((((...)))))..))))))))).((((((....)))))) ( -29.70) >DroSec_CAF1 1867 94 - 1 UGUGC---ACACACGCAUCAA----CUUCCCAUCCCCUUGGAGUACGAG-------------------UAUGAGCAACUGCAACAUUGUUGCCAUGCUCCUCGCUCCGGAGGUAUCCGGA .((((---......))))...----..(((.....(((((((((..(((-------------------((((.(((((.........))))))))))))...))))).)))).....))) ( -32.20) >DroSim_CAF1 1868 94 - 1 UGUGC---ACACACGCAUCAA----CUUCCCAUCCCCAUGGAGUACGAG-------------------UACGAGCAACUGCAACAUUGUUGCCAUGCUCCUCGCUCCGGAGGUAUCCGGA .((((---......))))..(----((..((((....))))))).((((-------------------...(((((...(((((...)))))..))))))))).((((((....)))))) ( -28.70) >DroEre_CAF1 2210 116 - 1 UGUGC---ACACACGCAUCCACAUGCUUCCAUUGCACUG-GAGUGCGAGCACAAGCACCCAGCACCAGCACGAACAACUGCAACAUUGUUGCCAUGCUCCUCGCUCCGGAGGUAUCCGGU .((((---......))))((..((((((((...((((..-..))))((((...((((....((....))..........(((((...)))))..))))....)))).))))))))..)). ( -34.10) >DroYak_CAF1 2239 91 - 1 UGUGCAGCACACACGUAUCUA----CUUCCAUUCCCCUU-GG------------------------AGCACGAACAACUGCAACAUUGUUGCCAUGCUGCUCGCUCCGGAGGUAUCCGGU .(.((((((.....((.((..----.(((((.......)-))------------------------))...))))....(((((...)))))..)))))).)...(((((....))))). ( -26.70) >consensus UGUGC___ACACACGCAUCAA____CUUCCCAUCCCCUUGGAGUACGAG___________________UACGAGCAACUGCAACAUUGUUGCCAUGCUCCUCGCUCCGGAGGUAUCCGGA ((((.....)))).((.............(((......)))..............................(((((...(((((...)))))..)))))...)).(((((....))))). (-18.70 = -19.50 + 0.80)

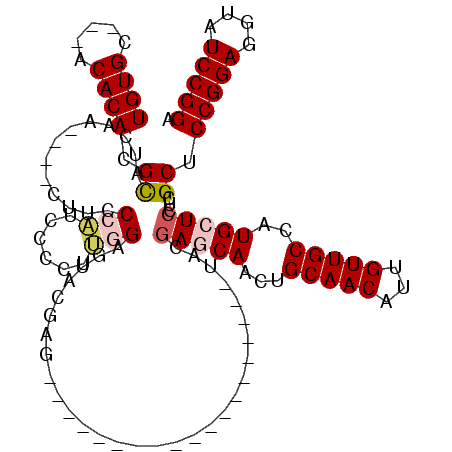
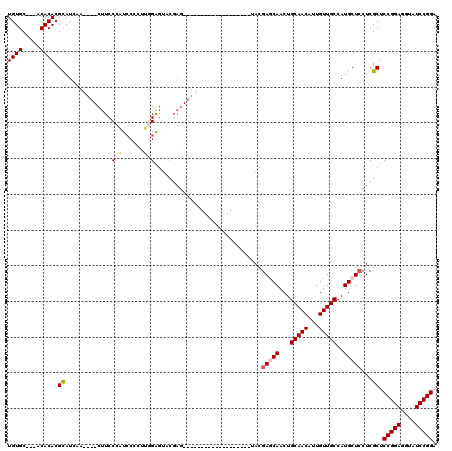
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:08 2006