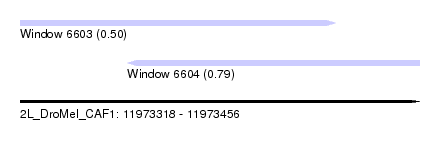
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,973,318 – 11,973,456 |
| Length | 138 |
| Max. P | 0.785889 |

| Location | 11,973,318 – 11,973,427 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 79.90 |
| Mean single sequence MFE | -25.22 |
| Consensus MFE | -13.61 |
| Energy contribution | -13.65 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.54 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
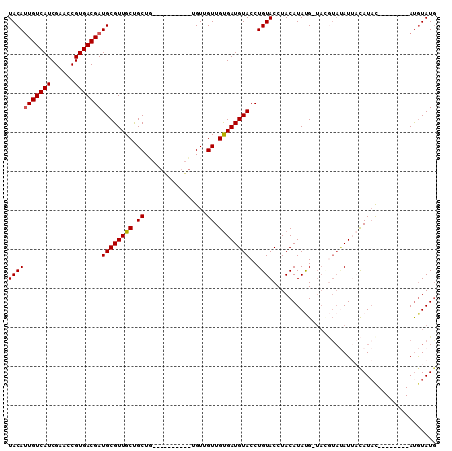
>2L_DroMel_CAF1 11973318 109 + 22407834 UACACUGUCAUCGAACCGUGACGAUGCGUUGCUGCUGCUGCUGUUGUUGUUGUUGUGAUGUACCUGUACCUACAUAUG-UAGGUAUAUUACAUACAUAUAUGUAUGUAUG ......((((((((((.(..((((((.((.((....)).))))))))..).))).))))).)).(((((((((....)-)))))))).((((((((....)))))))).. ( -34.60) >DroSec_CAF1 192839 89 + 1 UACAUUGUCAUCGAACCGUGACGAUGCGUUGCUGCU------------GUUGUUGUGAUGUACCUGUACCUACAUAUG-UACGUAUAUUACAUAU--------AUGUAUG ..(((((((((......)))))))))......((((------------((...((((((((((..((((........)-))))))))))))).))--------).))).. ( -23.00) >DroSim_CAF1 159222 89 + 1 UACAUUGUCAUCGAACCGUGACGAUGCGUUGCUGCU------------GUUGUUGUGAUGUACCUGUACCUACAUAUG-UACGUAUAUUACAUAC--------AUGUAUG ..(((((((((......)))))))))......((((------------((...((((((((((..((((........)-))))))))))))).))--------).))).. ( -24.90) >DroEre_CAF1 167261 86 + 1 UACAUUGUCAUCGAACCGUGACGAUGCGUUGCCGCUG---U------UGUUGUUGCGAUGUACCUGUACCUACAUACGA-------AUACUAUAU--------ACGUAUG (((((((((((......)))))))((((((((.((..---.------....)).))))))))..))))....((((((.-------(((...)))--------.)))))) ( -22.00) >DroYak_CAF1 164061 99 + 1 UACAUUGUCAUCGAACCGUGACGAUGCGUUGCUGCUG---UGGUUGCUGUUGUUGUGAUGUACCUGUACCUACAUACGAUAUAUACAUACCAUCC--------AUAUAUA ..(((((((((......)))))))))..........(---((((((.((((((((((.((((........))))))))))).))))).)))))..--------....... ( -21.60) >consensus UACAUUGUCAUCGAACCGUGACGAUGCGUUGCUGCUG__________UGUUGUUGUGAUGUACCUGUACCUACAUAUG_UACGUAUAUUACAUAC________AUGUAUG (((((((((((......)))))))((((((((.((................)).))))))))..)))).......................................... (-13.61 = -13.65 + 0.04)


| Location | 11,973,355 – 11,973,456 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 75.68 |
| Mean single sequence MFE | -24.42 |
| Consensus MFE | -12.49 |
| Energy contribution | -12.55 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.785889 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
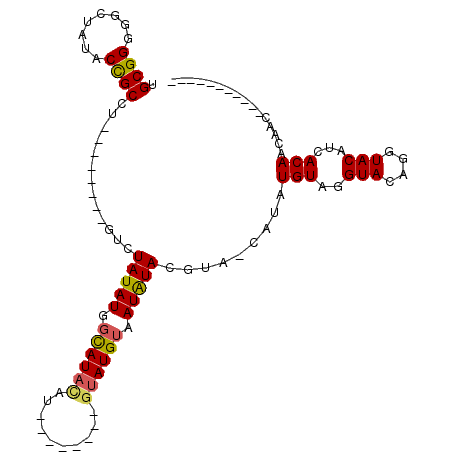
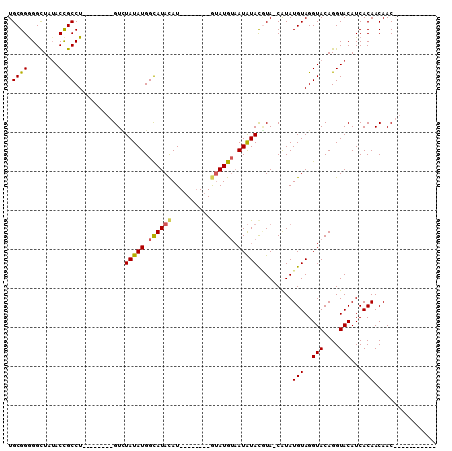
>2L_DroMel_CAF1 11973355 101 - 22407834 UGCGGGGGCUAUACCGCCU--------GUGUAUAUGGCAUACAUACAUAUAUGUAUGUAAUAUACCUA-CAUAUGUAGGUACAGGUACAUCACAACAACAACAACAGCAG (((((........))((((--------((((((((.(((((((((....))))))))).)))))((((-(....))))))))))))....................))). ( -32.30) >DroSec_CAF1 192875 82 - 1 UGCGGGGGCUAUACCGCCU--------GUCUAUAUGGCAUACAU--------AUAUGUAAUAUACGUA-CAUAUGUAGGUACAGGUACAUCACAACAAC----------- ((.((.(.((.(((((((.--------........)))....((--------(((((((.......))-))))))).)))).)).)...)).)).....----------- ( -18.60) >DroSim_CAF1 159258 82 - 1 UGCGGAGGCUAUACCGCCU--------GUUUAUAUGGCAUACAU--------GUAUGUAAUAUACGUA-CAUAUGUAGGUACAGGUACAUCACAACAAC----------- ....((.(((.(((((((.--------........))).(((((--------(((((((...))))))-))..))))))))..)))...))........----------- ( -19.70) >DroYak_CAF1 164098 99 - 1 UGCGGGGGUUAAACUGCCUGUUUGUCUGUCUAUAUGGUAUAUAU--------GGAUGGUAUGUAUAUAUCGUAUGUAGGUACAGGUACAUCACAACAACAGCAACCA--- (((.(..(((....(((((((...((((..((((((((((((((--------(.......))))))))))))))))))).)))))))......)))..).)))....--- ( -27.10) >consensus UGCGGGGGCUAUACCGCCU________GUCUAUAUGGCAUACAU________GUAUGUAAUAUACGUA_CAUAUGUAGGUACAGGUACAUCACAACAAC___________ .((((........)))).............(((((.((((((..........)))))).))))).........(((..(((....)))...)))................ (-12.49 = -12.55 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:42 2006