| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,922,649 – 11,922,743 |
| Length | 94 |
| Max. P | 0.999952 |

| Location | 11,922,649 – 11,922,743 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 97.44 |
| Mean single sequence MFE | -17.29 |
| Consensus MFE | -16.71 |
| Energy contribution | -16.91 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.18 |
| Structure conservation index | 0.97 |
| SVM decision value | 4.81 |
| SVM RNA-class probability | 0.999952 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11922649 94 + 22407834 CCUCCUUUUCUAGCCAGCCAGCGCUGACUCUUUCUAUUUUUUUGCUCCAUCUCUCUCUAUUACUUUGCCGCUGGCAUUUACAUGGACAAUCUAC ..(((.......((((((..(((..(....................................)..))).))))))........)))........ ( -12.28) >DroSec_CAF1 142078 93 + 1 CCUCCUUUUCUAGCCAGCCAGCGCUGGCUCUUUCUAUUU-UUUGCUCCAUCUCUCUCUAUUACUUUGCCGCUGGCAUUUACAUGGACAAUCUAC ...........(((((((....)))))))..........-.(((.(((((...............((((...)))).....))))))))..... ( -18.25) >DroSim_CAF1 108627 93 + 1 CCUCCUUUUCUAGCCAGCCAGCGCUGGCUCUUUCUAUUU-UUUGCUCCAUCUCUCUCUAUUACUUUGCCGCUGGCAUUUACAUGGACAAUCUAC ...........(((((((....)))))))..........-.(((.(((((...............((((...)))).....))))))))..... ( -18.25) >DroEre_CAF1 113263 93 + 1 CCUCCUUUUCUAGCCAGCCAGCGCUGGCUCUUUCUAUUU-UUUGCUCCAUCUCUCUCUAUUACUUUGCCACUGGCAUUUACAUGGACAAUCUAC ...........(((((((....)))))))..........-.(((.(((((...............((((...)))).....))))))))..... ( -18.25) >DroYak_CAF1 111253 94 + 1 CCUCCUUUUCUAGCCAGCCAGCGCUGGCUCUUUCUAUUUUUUUGCUCCAUCUCUCUCUGUUACAUUGCCACUGGCAUUUACAUGGACAAUCUAC ...........(((((((....)))))))............(((.((((........(((.....((((...))))...))))))))))..... ( -19.40) >consensus CCUCCUUUUCUAGCCAGCCAGCGCUGGCUCUUUCUAUUU_UUUGCUCCAUCUCUCUCUAUUACUUUGCCGCUGGCAUUUACAUGGACAAUCUAC ...........(((((((....)))))))............(((.(((((...............((((...)))).....))))))))..... (-16.71 = -16.91 + 0.20)

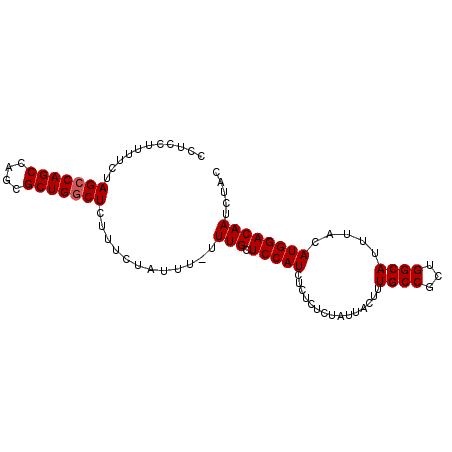
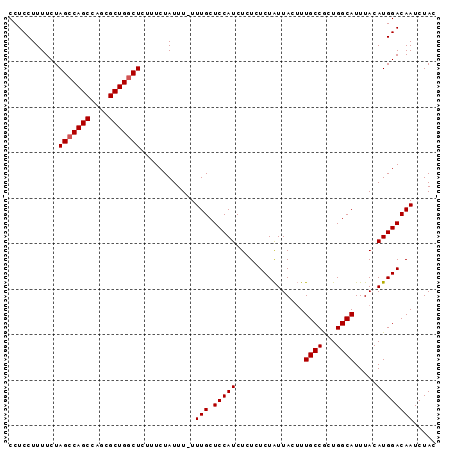
| Location | 11,922,649 – 11,922,743 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 97.44 |
| Mean single sequence MFE | -21.16 |
| Consensus MFE | -20.71 |
| Energy contribution | -20.55 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.03 |
| SVM RNA-class probability | 0.999763 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11922649 94 - 22407834 GUAGAUUGUCCAUGUAAAUGCCAGCGGCAAAGUAAUAGAGAGAGAUGGAGCAAAAAAAUAGAAAGAGUCAGCGCUGGCUGGCUAGAAAAGGAGG .....((((((((.(...((((...)))).(....)......).))))).)))............(((((((....)))))))........... ( -18.80) >DroSec_CAF1 142078 93 - 1 GUAGAUUGUCCAUGUAAAUGCCAGCGGCAAAGUAAUAGAGAGAGAUGGAGCAAA-AAAUAGAAAGAGCCAGCGCUGGCUGGCUAGAAAAGGAGG .....((((((((.(...((((...)))).(....)......).))))).))).-..........(((((((....)))))))........... ( -21.50) >DroSim_CAF1 108627 93 - 1 GUAGAUUGUCCAUGUAAAUGCCAGCGGCAAAGUAAUAGAGAGAGAUGGAGCAAA-AAAUAGAAAGAGCCAGCGCUGGCUGGCUAGAAAAGGAGG .....((((((((.(...((((...)))).(....)......).))))).))).-..........(((((((....)))))))........... ( -21.50) >DroEre_CAF1 113263 93 - 1 GUAGAUUGUCCAUGUAAAUGCCAGUGGCAAAGUAAUAGAGAGAGAUGGAGCAAA-AAAUAGAAAGAGCCAGCGCUGGCUGGCUAGAAAAGGAGG .....((((((((.(...((((...)))).(....)......).))))).))).-..........(((((((....)))))))........... ( -21.50) >DroYak_CAF1 111253 94 - 1 GUAGAUUGUCCAUGUAAAUGCCAGUGGCAAUGUAACAGAGAGAGAUGGAGCAAAAAAAUAGAAAGAGCCAGCGCUGGCUGGCUAGAAAAGGAGG ........(((.......((((...)))).(((..((........))..))).............(((((((....)))))))......))).. ( -22.50) >consensus GUAGAUUGUCCAUGUAAAUGCCAGCGGCAAAGUAAUAGAGAGAGAUGGAGCAAA_AAAUAGAAAGAGCCAGCGCUGGCUGGCUAGAAAAGGAGG .....((((((((.(...((((...)))).............).))))).)))............(((((((....)))))))........... (-20.71 = -20.55 + -0.16)

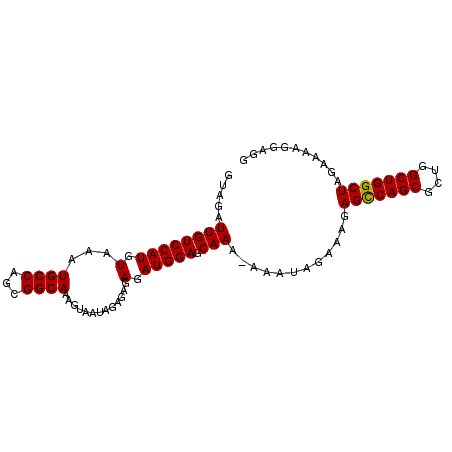
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:10:35 2006