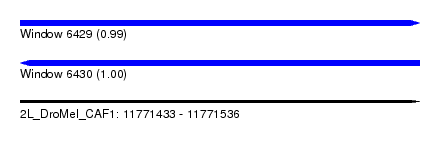

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,771,433 – 11,771,536 |
| Length | 103 |
| Max. P | 0.999669 |

| Location | 11,771,433 – 11,771,536 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 92.63 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -22.95 |
| Energy contribution | -23.15 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.12 |
| SVM RNA-class probability | 0.988500 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
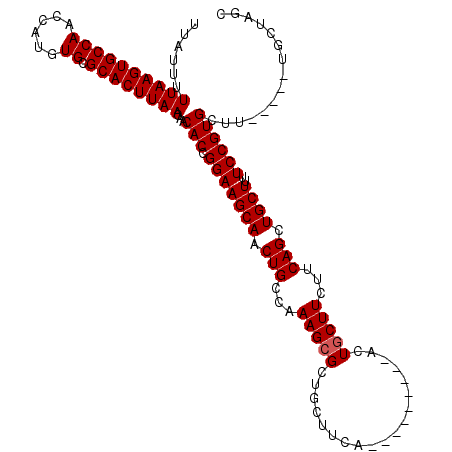
>2L_DroMel_CAF1 11771433 103 + 22407834 UUAUUUUUAAGUGCCAACCAUGUGCGCACUUAACAACACCGGAAGCAACUGCCAAAGAGCUGCUUCA---------ACUGCUUCUUCAGCUGCUUUUCCGUGCUU-----UGCUAGC ......((((((((((......)).))))))))(((..(((((((((.(((...((((((.......---------...)).))))))).))))..)))).)..)-----))..... ( -25.30) >DroSec_CAF1 29439 103 + 1 UUAUUUUUAAGUGCCAACCAUGUGCGCACUUAACAACACCGGAAGCAACUGCCAAAGCGCUGCUUCC---------ACUGCUUCUUCAGCUGCUUUUCCGUGCUU-----UGCUAGC ......((((((((((......)).))))))))(((..(((((((((.(((...(((((.((....)---------).)))))...))).))))..)))).)..)-----))..... ( -27.00) >DroSim_CAF1 28555 103 + 1 UUAUUUUUAAGUGCCAACCAUGUGCGCACUUAACAACACCGGAAGCAACUGCCAAAGCGCUGCUUCC---------ACUGCUUCUUCAGCUGCUUUUCCGUGCUU-----UCCUAGC ......((((((((((......)).))))))))......((((((((.(((...(((((.((....)---------).)))))...))).))))..)))).(((.-----....))) ( -26.80) >DroEre_CAF1 39919 112 + 1 UUAUUUUUAAGUGCCAACCAUGUGUGCACUUAACAACACCGGAAGCAACUGCCAAAGCGCUGCUUCAACUGCUUCAACUGCUUCUUCAGCUGCUUUUCCGUGCUU-----UGCGAGC ......((((((((((......)).))))))))(((..(((((((((.(((...(((((..((.......))......)))))...))).))))..)))).)..)-----))..... ( -26.70) >DroYak_CAF1 30829 108 + 1 UUAUUUUUAAGUGCCAACCAUGUGUGCACUUAACAACACCGGAAGCAACUGCCAAAGCGCUGCUUCA---------ACUGCUUCUUCAGCUGCUUUUCCGUGCUUUGCUUUGCUAGC ......((((((((((......)).))))))))..........(((((..((.(((((((.......---------...(((.....))).........))))))))).)))))... ( -26.41) >consensus UUAUUUUUAAGUGCCAACCAUGUGCGCACUUAACAACACCGGAAGCAACUGCCAAAGCGCUGCUUCA_________ACUGCUUCUUCAGCUGCUUUUCCGUGCUU_____UGCUAGC ......((((((((((......)).))))))))...(((.(((((((.(((...(((((...................)))))...))).))))..))))))............... (-22.95 = -23.15 + 0.20)

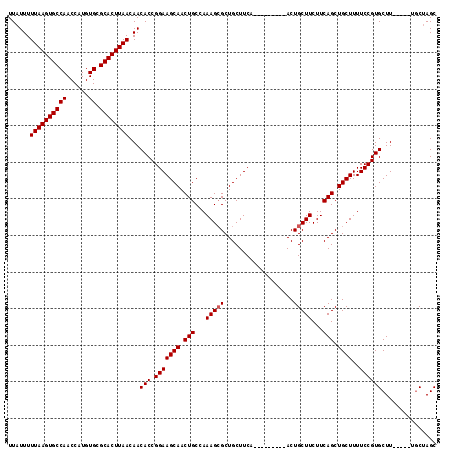
| Location | 11,771,433 – 11,771,536 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 92.63 |
| Mean single sequence MFE | -40.68 |
| Consensus MFE | -34.56 |
| Energy contribution | -34.76 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.55 |
| Structure conservation index | 0.85 |
| SVM decision value | 3.86 |
| SVM RNA-class probability | 0.999669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11771433 103 - 22407834 GCUAGCA-----AAGCACGGAAAAGCAGCUGAAGAAGCAGU---------UGAAGCAGCUCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCGCACAUGGUUGGCACUUAAAAAUAA (((....-----.))).((((..((((((((..((((.(((---------((...)))))))))..)))))))))))).((((.((((((((.((......)))))))))).)))). ( -39.30) >DroSec_CAF1 29439 103 - 1 GCUAGCA-----AAGCACGGAAAAGCAGCUGAAGAAGCAGU---------GGAAGCAGCGCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCGCACAUGGUUGGCACUUAAAAAUAA (((....-----.))).((((..((((((((..(((((..(---------(....))..)))))..)))))))))))).((((.((((((((.((......)))))))))).)))). ( -40.20) >DroSim_CAF1 28555 103 - 1 GCUAGGA-----AAGCACGGAAAAGCAGCUGAAGAAGCAGU---------GGAAGCAGCGCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCGCACAUGGUUGGCACUUAAAAAUAA (((....-----.))).((((..((((((((..(((((..(---------(....))..)))))..)))))))))))).((((.((((((((.((......)))))))))).)))). ( -40.20) >DroEre_CAF1 39919 112 - 1 GCUCGCA-----AAGCACGGAAAAGCAGCUGAAGAAGCAGUUGAAGCAGUUGAAGCAGCGCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCACACAUGGUUGGCACUUAAAAAUAA (((....-----.))).((((..((((((((..(((((.((((...(....)...)))))))))..)))))))))))).((((.((((((((.((......)))))))))).)))). ( -42.40) >DroYak_CAF1 30829 108 - 1 GCUAGCAAAGCAAAGCACGGAAAAGCAGCUGAAGAAGCAGU---------UGAAGCAGCGCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCACACAUGGUUGGCACUUAAAAAUAA (((.....)))..((((((((..((((((((..(((((.((---------((...)))))))))..))))))))))).))))).((((((((.((......))))))))))...... ( -41.30) >consensus GCUAGCA_____AAGCACGGAAAAGCAGCUGAAGAAGCAGU_________UGAAGCAGCGCUUUGGCAGUUGCUUCCGGUGUUGUUAAGUGCGCACAUGGUUGGCACUUAAAAAUAA (((..........))).((((..((((((((..(((((.....................)))))..)))))))))))).((((.((((((((.((......)))))))))).)))). (-34.56 = -34.76 + 0.20)

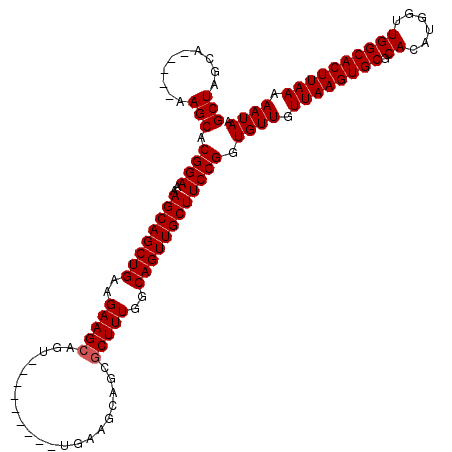

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:08:50 2006