
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,617,214 – 11,617,340 |
| Length | 126 |
| Max. P | 0.997184 |

| Location | 11,617,214 – 11,617,320 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 78.59 |
| Mean single sequence MFE | -32.97 |
| Consensus MFE | -19.24 |
| Energy contribution | -20.00 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.667524 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11617214 106 - 22407834 AGAAGAGCCA--GGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUUGCGGCGAGAUAAAGAGUCCAUGGACAGAGUUGGU------AAUUCCGCAGCAUAA ...(((((..--(((((....))))).....)))))((((((((((((((....))))))))))..........(((....))).........------....))))....... ( -35.30) >DroSec_CAF1 70512 106 - 1 AGAAGAGCCA--GGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUGGCGGCGAGAUAAAGAGUCCAUGGACAGAGUUGGU------AAUUCCGCAGCAUAA ...(((((..--(((((....))))).....)))))(((((((((.((((....)))).)))))..........(((....))).........------....))))....... ( -31.20) >DroSim_CAF1 74297 106 - 1 AGAAGAGCCA--GGACUUUUGGGUUCGUUAAGCUCAGCGGGCUGCAAUAUGGGUGUGUUGUGGCGAGAUAAAGAGUCCAUGGACAGAGUUGGU------AAUGCCGCAGCAUAA ......((((--((((((((...((((((.((((.....))))(((((((....)))))))))))))...)))))))).))).).........------.((((....)))).. ( -32.50) >DroEre_CAF1 100990 104 - 1 AGCAGAGCCA--GGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAA--UGGGUGUGUUGCGGGGAGAUGCAGAGUCCAUGGACAAAUUUGCU------GAUUCCGCAGCAUAA .(((((((..--(((((....))))).....))))))).........--....((((((((((((....((((((((....)))...))))).------..)))))))))))). ( -41.10) >DroYak_CAF1 73720 106 - 1 AGAAGAGCCA--GGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGCUGUGUGGCGGGGAGAUAAAGAGUCCAUGGAGAGAUUUGUU------AAUUCCACGGCAUAA ......(((.--((((((((...(((.(...(((..((((.(........).))))..))).).)))...)))))))).(((((..(.....)------..))))).))).... ( -31.80) >DroAna_CAF1 83667 107 - 1 AAAAGUGGCUGUGGGGCUUUGGGUUCGUUAAGCUUUGCGGGUUAAAAUGAAAGAUAUUUAUAAAGAUUUAGUGAGUCC-----CAGAGGUAACCAAGUGAAUUUUGC--CAGGA .....((((..(((..(((((((((((((((((((((..(((.............)))..))))).))))))))).))-----)))))....)))..........))--))... ( -25.92) >consensus AGAAGAGCCA__GGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUUGCGGCGAGAUAAAGAGUCCAUGGACAGAGUUGGU______AAUUCCGCAGCAUAA ...(((((....(((((....))))).....)))))((((.(((((((((....)))))))))...........(((....)))...................))))....... (-19.24 = -20.00 + 0.76)


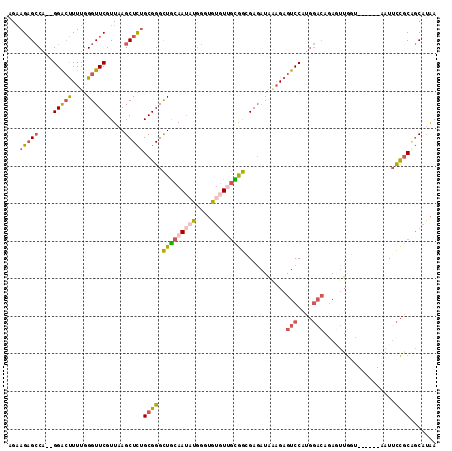
| Location | 11,617,242 – 11,617,340 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 93.56 |
| Mean single sequence MFE | -28.28 |
| Consensus MFE | -24.40 |
| Energy contribution | -25.84 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.81 |
| SVM RNA-class probability | 0.997184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11617242 98 - 22407834 AUAUUUAU--AUAAAAGUAAAUAGAAGAGCCAGGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUUGCGGCGAGAUAAAGAGUCCAU .(((((((--......))))))).........((((((((...((((((.((((.....))))(((((((....)))))))))))))...)))))))).. ( -31.60) >DroSec_CAF1 70540 100 - 1 AUAUUUAUAUAUAAAAGUAAAUAGAAGAGCCAGGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUGGCGGCGAGAUAAAGAGUCCAU .(((((((........))))))).........((((((((...((((((.((((.....))))((.((((....)))).))))))))...)))))))).. ( -27.30) >DroSim_CAF1 74325 100 - 1 AUAUUUAUAUAUAAAAGUAAAUAGAAGAGCCAGGACUUUUGGGUUCGUUAAGCUCAGCGGGCUGCAAUAUGGGUGUGUUGUGGCGAGAUAAAGAGUCCAU .(((((((........))))))).........((((((((...((((((.((((.....))))(((((((....)))))))))))))...)))))))).. ( -29.10) >DroEre_CAF1 101018 96 - 1 AUAUUUAU--AUAAAAGUAAAUAGCAGAGCCAGGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAA--UGGGUGUGUUGCGGGGAGAUGCAGAGUCCAU .(((((((--......)))))))(((((((..(((((....))))).....)))))))((((((((.--....))).(((((......)))))))))).. ( -30.10) >DroYak_CAF1 73748 98 - 1 AUAUUCAU--GUAAAAGUAAAUAGAAGAGCCAGGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGCUGUGUGGCGGGGAGAUAAAGAGUCCAU ........--......................((((((((...(((.(...(((..((((.(........).))))..))).).)))...)))))))).. ( -23.30) >consensus AUAUUUAU__AUAAAAGUAAAUAGAAGAGCCAGGACUUUUGGGUUCGUUAAGCUCUGCGGGCUGCAAUAUGGGUGUGUUGCGGCGAGAUAAAGAGUCCAU .(((((((........))))))).........((((((((...((((((.((((.....))))(((((((....)))))))))))))...)))))))).. (-24.40 = -25.84 + 1.44)


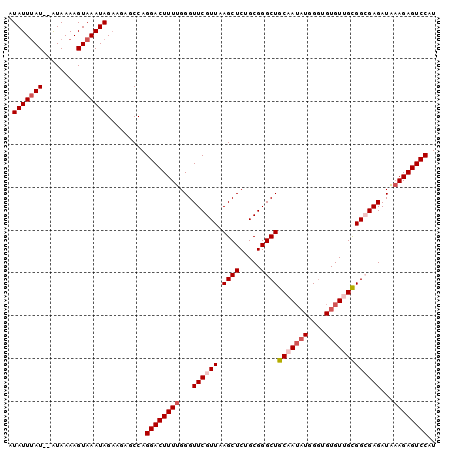
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:35 2006