
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,600,775 – 11,600,865 |
| Length | 90 |
| Max. P | 0.999872 |

| Location | 11,600,775 – 11,600,865 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 89.45 |
| Mean single sequence MFE | -22.42 |
| Consensus MFE | -18.42 |
| Energy contribution | -19.08 |
| Covariance contribution | 0.66 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11600775 90 + 22407834 CA----UAUAUA------UGUGUGUAUAUUUAUGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGUGCUUAUAAAUUGGGUGGUUGAAAGGUUGCG-- ((----((.(((------((....))))).))))...((((((((((((.((((((.((((((.....)))))).))))))...))))))...)))))).-- ( -21.10) >DroSec_CAF1 55033 98 + 1 CA----UAUAUAUUUAUGUGUGUGUAUAUUUAUGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGUGCUUAUAAAUUGGGUGGAUGAAAGGUUGCGGU ((----((((((....)))))))).........(((....(((((.((..((((((.((((((.....)))))).)))))))).)))))....)))...... ( -23.20) >DroSim_CAF1 57684 102 + 1 CACAUAUAUAUAUAUAUGUGUGUGUAUAUUUAUGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGUGCUUAUAAAUUGGGUGGAUGAAAGGUUGCGGU ((((((((((....)))))))))).........(((....(((((.((..((((((.((((((.....)))))).)))))))).)))))....)))...... ( -27.50) >DroEre_CAF1 85885 88 + 1 CA----UAU--A--------UAUGUACAUUUAUGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGUGCUUAUAAAUUGGGUGGAUGAAAGGUUGCGGU ..----...--.--------......((((((((((((....)))))...((((((.((((((.....)))))).))))))..)))))))............ ( -20.30) >DroYak_CAF1 56680 90 + 1 CA----UAUAUA--------UAUGUAUAUUUACGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGCGCUUAUAAAUUGGGUGGAUGAAAGGUUGCGGU ((----...(((--------(.((((.((((..(((((....)))))..)))).)))).))))..))..((((((.....))))))................ ( -20.00) >consensus CA____UAUAUA______UGUGUGUAUAUUUAUGCCGGCAAUCCGGCUGAAAUUUACAAGUAUUUUGGGUGCUUAUAAAUUGGGUGGAUGAAAGGUUGCGGU ......(((((((......)))))))((((((((((((....)))))...((((((.((((((.....)))))).))))))..)))))))............ (-18.42 = -19.08 + 0.66)

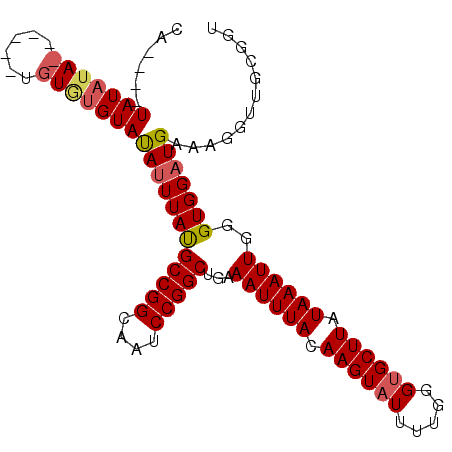
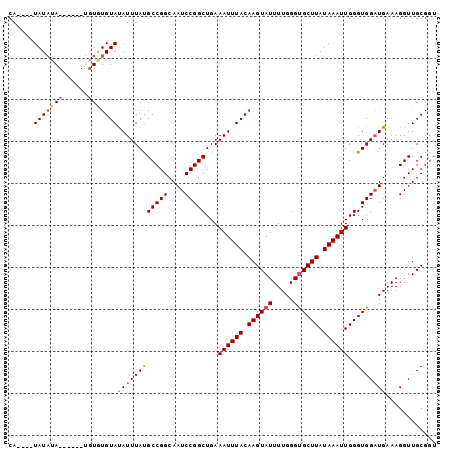
| Location | 11,600,775 – 11,600,865 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 89.45 |
| Mean single sequence MFE | -16.48 |
| Consensus MFE | -14.90 |
| Energy contribution | -14.90 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.33 |
| SVM RNA-class probability | 0.999872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11600775 90 - 22407834 --CGCAACCUUUCAACCACCCAAUUUAUAAGCACCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCAUAAAUAUACACACA------UAUAUA----UG --...................((((((((((...........))))))))))...(((((....)))))...............------......----.. ( -14.90) >DroSec_CAF1 55033 98 - 1 ACCGCAACCUUUCAUCCACCCAAUUUAUAAGCACCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCAUAAAUAUACACACACAUAAAUAUAUA----UG .....................((((((((((...........))))))))))...(((((....)))))...........................----.. ( -14.90) >DroSim_CAF1 57684 102 - 1 ACCGCAACCUUUCAUCCACCCAAUUUAUAAGCACCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCAUAAAUAUACACACACAUAUAUAUAUAUAUGUG .....................((((((((((...........))))))))))...(((((....))))).............((((((((....)))))))) ( -21.30) >DroEre_CAF1 85885 88 - 1 ACCGCAACCUUUCAUCCACCCAAUUUAUAAGCACCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCAUAAAUGUACAUA--------U--AUA----UG ...(((...............((((((((((...........))))))))))...(((((....))))).....))).....--------.--...----.. ( -16.00) >DroYak_CAF1 56680 90 - 1 ACCGCAACCUUUCAUCCACCCAAUUUAUAAGCGCCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCGUAAAUAUACAUA--------UAUAUA----UG ...((................((((((((((...........))))))))))...(((((....)))))))...........--------......----.. ( -15.30) >consensus ACCGCAACCUUUCAUCCACCCAAUUUAUAAGCACCCAAAAUACUUGUAAAUUUCAGCCGGAUUGCCGGCAUAAAUAUACACACA______UAUAUA____UG .....................((((((((((...........))))))))))...(((((....)))))................................. (-14.90 = -14.90 + 0.00)

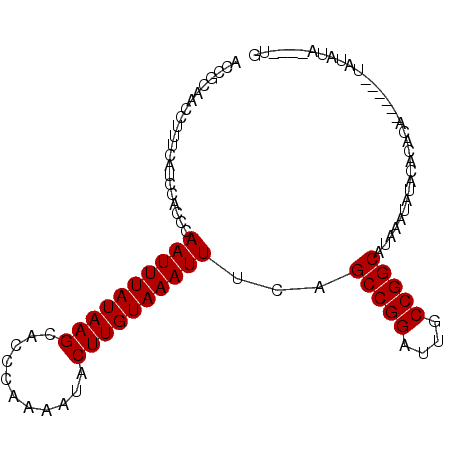
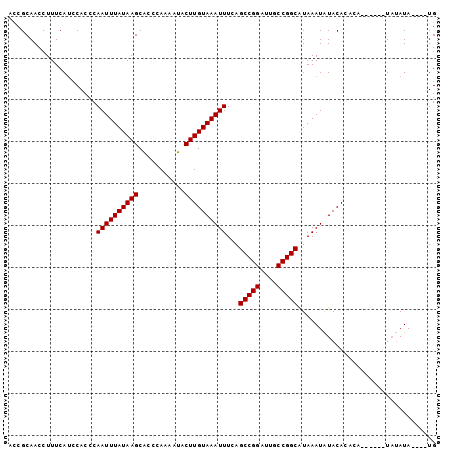
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:26 2006