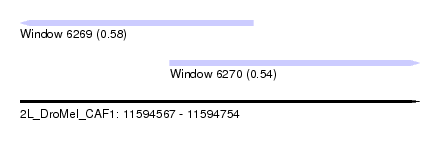
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,594,567 – 11,594,754 |
| Length | 187 |
| Max. P | 0.575645 |

| Location | 11,594,567 – 11,594,676 |
|---|---|
| Length | 109 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 89.27 |
| Mean single sequence MFE | -25.18 |
| Consensus MFE | -17.79 |
| Energy contribution | -17.73 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.575645 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
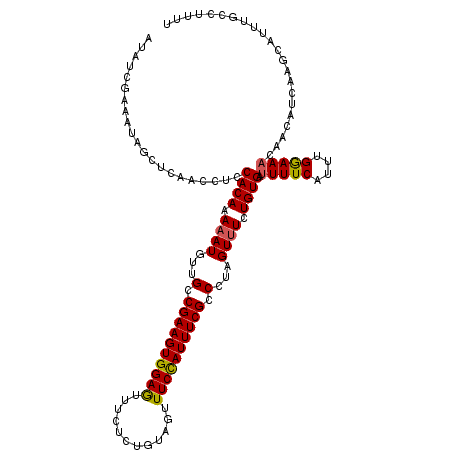
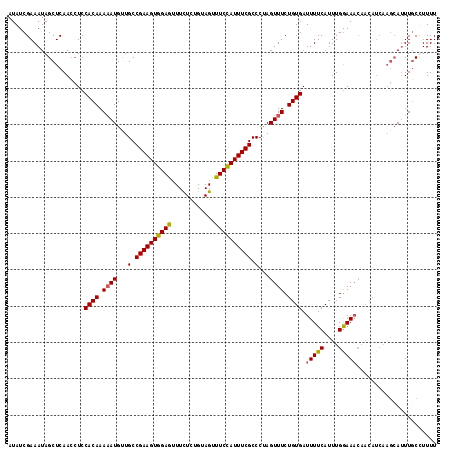
>2L_DroMel_CAF1 11594567 109 - 22407834 AUAUCGAAAUAGCUCAACAUCCACAAAUAUGUUGCCGAAGUGGAAUUUCUCUGUAGUUUCCAUUUCGCCCUAGUUUCUGUGAUUUUCAUUUGGAAUCU---------AGUUGCUUUUU ......(((.(((.((((((........)))))).((((((((((...((....)).))))))))))..((((.(((((((.....)))..)))).))---------))..))).))) ( -23.80) >DroSec_CAF1 48685 118 - 1 AUAUCGAAAUAGCUCAACCUCCACAAAAAUGUUGCCGAAGUGGAGUUUCUCUGUAGUUUCCAUUUCGCCCUAGUUUCUGUGAUUUUCAUUUGGAAACAACAUCAAGCAUUUGCUUUUU .....((..............((((.((((...(.((((((((((...((....)).)))))))))).)...)))).))))........(((....)))..))((((....))))... ( -23.80) >DroSim_CAF1 51398 118 - 1 AUAUCGAAAUAGCUCAAGCUCCACAAAAAUGUUGCCGAAGUGGAGUUUCUCUGUAGUUUCCAUUUCGCCCUAGUUUCUGUGAUUUUCAUUUGGAAACAACAUCAAGCAUUUCCCUUUU .....(((((.(((.......((((.((((...(.((((((((((...((....)).)))))))))).)...)))).))))........(((....))).....))))))))...... ( -26.40) >DroYak_CAF1 50584 118 - 1 AUAUCGAAAUAGCUCAACCUCCACAAAAAUGCUGCCGAAGUGGAGUUUUUGUGUAGUUUCUAUUUCGCCCCAGUUUCUGUGAUUUUCAUUUGAAAAGAACAUCAAGCAUUUGCCUUUU ....((((((((...(((...(((((((((.(..(....)..).)))))))))..))).)))))))).....((((.(((..(((((....)))))..)))..))))........... ( -26.70) >consensus AUAUCGAAAUAGCUCAACCUCCACAAAAAUGUUGCCGAAGUGGAGUUUCUCUGUAGUUUCCAUUUCGCCCUAGUUUCUGUGAUUUUCAUUUGGAAACAACAUCAAGCAUUUGCCUUUU .....................((((.((((...(.((((((((((............)))))))))).)...)))).)))).(((((....)))))...................... (-17.79 = -17.73 + -0.06)

| Location | 11,594,637 – 11,594,754 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 91.82 |
| Mean single sequence MFE | -29.60 |
| Consensus MFE | -23.86 |
| Energy contribution | -24.05 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.540569 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
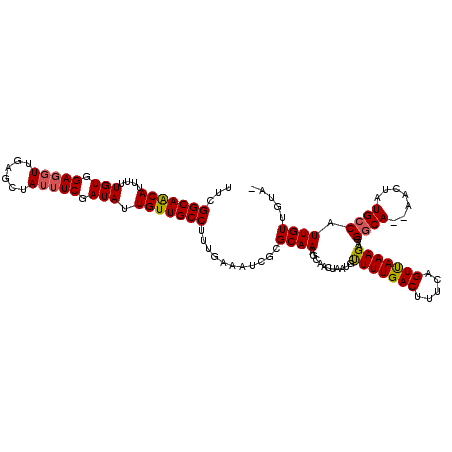
>2L_DroMel_CAF1 11594637 117 + 22407834 UUCGGCAACAUAUUUGUGGAUGUUGAGCUAUUUCGAUAUUGUUGCCUUUGAAAUCGCGCAAACCAACUAAUGUUUUGACUUUCAGUCAAAGAAGGGCA--AACUAUGCCAUUGUUGAAC ((((((((.(((((..(((...(((.((.(((((((...........))))))).)).))).)))...)))))((((((.....))))))....((((--.....)))).)))))))). ( -29.70) >DroSec_CAF1 48764 118 + 1 UUCGGCAACAUUUUUGUGGAGGUUGAGCUAUUUCGAUAUUGUUGCCUUUGAAAUCGCGCAAAGCAAAUAAUGUUUUGACUUUCAGUUAAAGAAGGGCAAAAACUAUGCCAUUGUUGUA- ....(((((((((((((((((((...((.(((((((...........))))))).)).(((((((.....))))))))))))))..))))))..((((.......))))..)))))).- ( -29.50) >DroSim_CAF1 51477 118 + 1 UUCGGCAACAUUUUUGUGGAGCUUGAGCUAUUUCGAUAUUGUUGCCUUUGAAAUAGCGCAAAGCAACUAAUGUUUUGACUUUCAGUUAAAGAAGGGCAAAAACUAUGCCAUUGUUGUA- ....((((((.....((((.((((..((((((((((...........))))))))))...)))).......(((((..(((((.......)))))...)))))....)))))))))).- ( -32.60) >DroYak_CAF1 50663 113 + 1 UUCGGCAGCAUUUUUGUGGAGGUUGAGCUAUUUCGAUAUUGUUGCCUUUGAAACCGCGCAA---AACUAAUGCUUUGACUUUCAGUUAAAGAAGGACA--AACUAUGCCAUUGUUGUG- ...(((((((....(((.(((((......))))).))).)))))))........(((((((---........(((((((.....)))))))..((.((--.....)))).))).))))- ( -26.60) >consensus UUCGGCAACAUUUUUGUGGAGGUUGAGCUAUUUCGAUAUUGUUGCCUUUGAAAUCGCGCAAAGCAACUAAUGUUUUGACUUUCAGUUAAAGAAGGGCA__AACUAUGCCAUUGUUGUA_ ...(((((((....(((.(((((......))))).))).)))))))...........((((...........(((((((.....)))))))...((((.......)))).))))..... (-23.86 = -24.05 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:14 2006