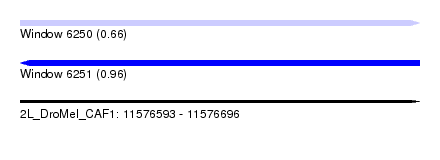

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,576,593 – 11,576,696 |
| Length | 103 |
| Max. P | 0.956859 |

| Location | 11,576,593 – 11,576,696 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 88.52 |
| Mean single sequence MFE | -30.93 |
| Consensus MFE | -24.58 |
| Energy contribution | -24.03 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.656669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
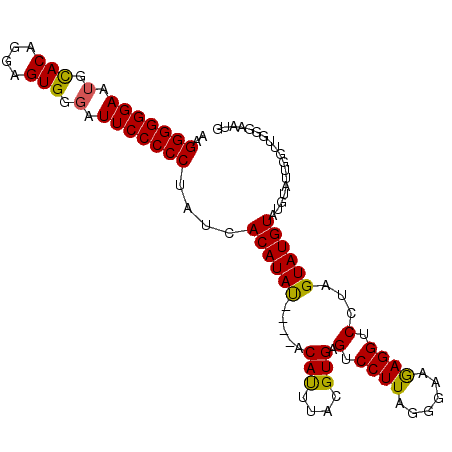
>2L_DroMel_CAF1 11576593 103 + 22407834 AAGGGGGGAAUGUACAGGAGUGGGAUUCCCCCAAUCACAUAUGUAUGCACUUACGUGAGUCCUUAGAAAUGAGGUCCUUGUAUGUAUGUAUUGGUUGGGAAUG ..(((((((.(.(((....))).).)))))))(((((.(((((((((((...((....))(((((....)))))....)))))))))))..)))))....... ( -32.40) >DroSec_CAF1 31047 99 + 1 AAGGGGGGAAUGCACAGGAGUGGGAUUCCCCCUAUCACAUAC----ACAUUUAUGUGAGUCCUUAGGGAAGAGGUCCUAGUAUGUAUGUAUUGGUUGGGAAUG .((((((((.(.(((....))).).)))))))).(((((((.----.....)))))))(.((((......)))).)(((((((....)))))))......... ( -32.00) >DroSim_CAF1 32731 99 + 1 AAGGGGGGAAAGCACAGGAGUGGGAUUCCCCCUAUCACAUAU----ACAUUUACGUGAGUCCUUACGGAGAAGGUCCUAGUAUGUAUGUAUUGGUUGGGAAUG ..((((((((..(((....)))...)))))))).((((.((.----.....)).)))).((((.(((((.....)))((((((....)))))))).))))... ( -28.40) >consensus AAGGGGGGAAUGCACAGGAGUGGGAUUCCCCCUAUCACAUAU____ACAUUUACGUGAGUCCUUAGGGAAGAGGUCCUAGUAUGUAUGUAUUGGUUGGGAAUG ..(((((((.(.(((....))).).)))))))....((((((.....(((....))).(.((((......)))).)...)))))).................. (-24.58 = -24.03 + -0.55)


| Location | 11,576,593 – 11,576,696 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 88.52 |
| Mean single sequence MFE | -25.83 |
| Consensus MFE | -20.82 |
| Energy contribution | -20.93 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.77 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.956859 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
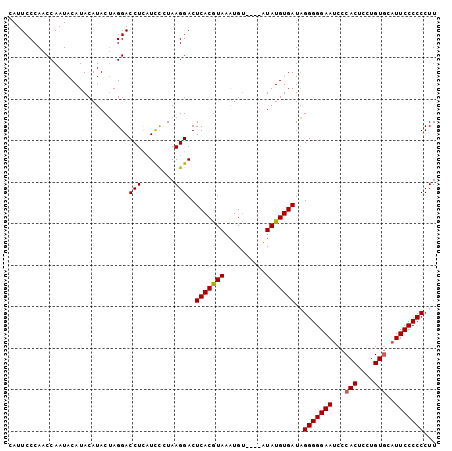
>2L_DroMel_CAF1 11576593 103 - 22407834 CAUUCCCAACCAAUACAUACAUACAAGGACCUCAUUUCUAAGGACUCACGUAAGUGCAUACAUAUGUGAUUGGGGGAAUCCCACUCCUGUACAUUCCCCCCUU .................((((((......(((........)))...(((....)))......))))))...((((((((...((....))..))))))))... ( -22.30) >DroSec_CAF1 31047 99 - 1 CAUUCCCAACCAAUACAUACAUACUAGGACCUCUUCCCUAAGGACUCACAUAAAUGU----GUAUGUGAUAGGGGGAAUCCCACUCCUGUGCAUUCCCCCCUU .............((((((((((.((((........)))).(......).....)))----)))))))...((((((((..(((....))).))))))))... ( -28.60) >DroSim_CAF1 32731 99 - 1 CAUUCCCAACCAAUACAUACAUACUAGGACCUUCUCCGUAAGGACUCACGUAAAUGU----AUAUGUGAUAGGGGGAAUCCCACUCCUGUGCUUUCCCCCCUU ...............((((.((((....((....(((....))).....))....))----)).))))...(((((((...(((....)))..)))))))... ( -26.60) >consensus CAUUCCCAACCAAUACAUACAUACUAGGACCUCAUCCCUAAGGACUCACGUAAAUGU____AUAUGUGAUAGGGGGAAUCCCACUCCUGUGCAUUCCCCCCUU .............................(((........)))..(((((((..........)))))))..(((((((...(((....)))..)))))))... (-20.82 = -20.93 + 0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:05:56 2006