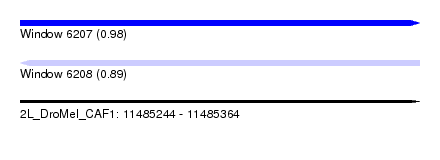

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,485,244 – 11,485,364 |
| Length | 120 |
| Max. P | 0.980452 |

| Location | 11,485,244 – 11,485,364 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -38.23 |
| Consensus MFE | -31.30 |
| Energy contribution | -32.17 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11485244 120 + 22407834 CAUCACCCACACGGAGGUCCACCUCCCCCUCGUCACCAUCAUCACCAUCGUGGCAGACAUGGUGCUGGUGGACGCUGUGGACACGGACACCAGCACCAGCCUUUUCCACCACCGCCUCAU ............(((((....))))).......................((((.(((..(((((((((((..((.((....))))..)))))))))))...))).))))........... ( -42.10) >DroSec_CAF1 49066 120 + 1 CAUCACCCACACGGAGGUCCGCCACCCCCUCGUCACCAUCACCACCAUCGUGGCAGACAUGGUGGUGGUAGACACUGUGGACACGGACCCCAGCACCAGCCUUUUCCACCACCCCCUCAU ............((.((((((((((......(((......((((((((((((.....)))))))))))).)))...))))...)))))))).((....)).................... ( -38.20) >DroSim_CAF1 52848 120 + 1 CAUCACCCACACGGAGGUCCGCCUCCCCCUCGUCACCAUCACCACCAUCGUGGCAGACAUGGUGGUGGUAGACACUGUGGACACGGACCCCAGCACCAGCCUUUUCCACCACCCCAUCAU ......(((((.(((((....))))).....(((......((((((((((((.....)))))))))))).)))..)))))....(((.....((....))....)))............. ( -39.80) >DroYak_CAF1 54726 114 + 1 CACCACCCGCCUGGAGGUCCGCCUCCCCCUCGCCAUCACCACCACCAUCGUGGCAGACAUGGUAGUGGUAGACACUGUGGACACGGAC------ACCAGCGUUUUCCACCACCCCAUCAU .......(((.(((..(((((..(((....((...(((((((.(((((.((.....))))))).))))).))...)).)))..)))))------.))))))................... ( -32.80) >consensus CAUCACCCACACGGAGGUCCGCCUCCCCCUCGUCACCAUCACCACCAUCGUGGCAGACAUGGUGGUGGUAGACACUGUGGACACGGACCCCAGCACCAGCCUUUUCCACCACCCCAUCAU ......((((..(((((....)))))..............((((((((((((.....)))))))))))).......))))....(((.....((....))....)))............. (-31.30 = -32.17 + 0.88)


| Location | 11,485,244 – 11,485,364 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -50.65 |
| Consensus MFE | -41.09 |
| Energy contribution | -41.52 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.888245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11485244 120 - 22407834 AUGAGGCGGUGGUGGAAAAGGCUGGUGCUGGUGUCCGUGUCCACAGCGUCCACCAGCACCAUGUCUGCCACGAUGGUGAUGAUGGUGACGAGGGGGAGGUGGACCUCCGUGUGGGUGAUG .....((.((.((((...(((((((((((((((..(((.......)))..))))))))))).)))).)))).)).))...........(.(.(.(((((....))))).).).)...... ( -54.60) >DroSec_CAF1 49066 120 - 1 AUGAGGGGGUGGUGGAAAAGGCUGGUGCUGGGGUCCGUGUCCACAGUGUCUACCACCACCAUGUCUGCCACGAUGGUGGUGAUGGUGACGAGGGGGUGGCGGACCUCCGUGUGGGUGAUG ....................(((.(..(.(((((((((..((.(..((((.((((.(((((((((......))).)))))).))))))))..).))..))))))))).)..).))).... ( -47.60) >DroSim_CAF1 52848 120 - 1 AUGAUGGGGUGGUGGAAAAGGCUGGUGCUGGGGUCCGUGUCCACAGUGUCUACCACCACCAUGUCUGCCACGAUGGUGGUGAUGGUGACGAGGGGGAGGCGGACCUCCGUGUGGGUGAUG ....................(((.(..(.(((((((((.(((.(..((((.((((.(((((((((......))).)))))).))))))))..).))).))))))))).)..).))).... ( -51.90) >DroYak_CAF1 54726 114 - 1 AUGAUGGGGUGGUGGAAAACGCUGGU------GUCCGUGUCCACAGUGUCUACCACUACCAUGUCUGCCACGAUGGUGGUGGUGAUGGCGAGGGGGAGGCGGACCUCCAGGCGGGUGGUG ...................((((((.------((((((.(((.(..((((((((((((((((((.....)).))))))))))))..))))..).))).))))))..))).)))....... ( -48.50) >consensus AUGAGGGGGUGGUGGAAAAGGCUGGUGCUGGGGUCCGUGUCCACAGUGUCUACCACCACCAUGUCUGCCACGAUGGUGGUGAUGGUGACGAGGGGGAGGCGGACCUCCGUGUGGGUGAUG ....................(((.(..(.(((((((((.(((.(..(((((..(((((((((((.....)).)))))))))...).))))..).))).))))))))).)..).))).... (-41.09 = -41.52 + 0.44)
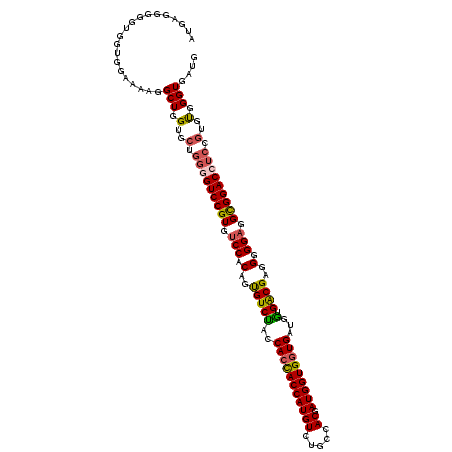



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:05:13 2006