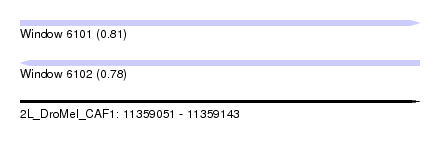

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,359,051 – 11,359,143 |
| Length | 92 |
| Max. P | 0.811297 |

| Location | 11,359,051 – 11,359,143 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 84.76 |
| Mean single sequence MFE | -27.60 |
| Consensus MFE | -17.68 |
| Energy contribution | -18.24 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.811297 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
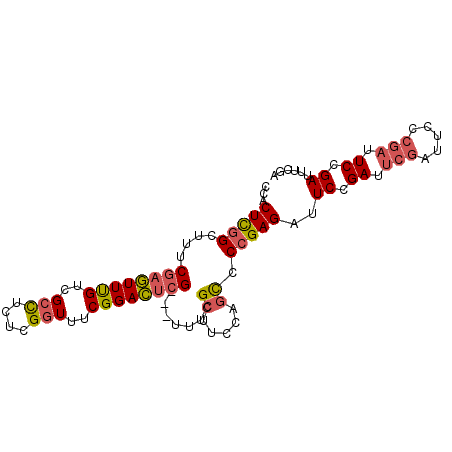
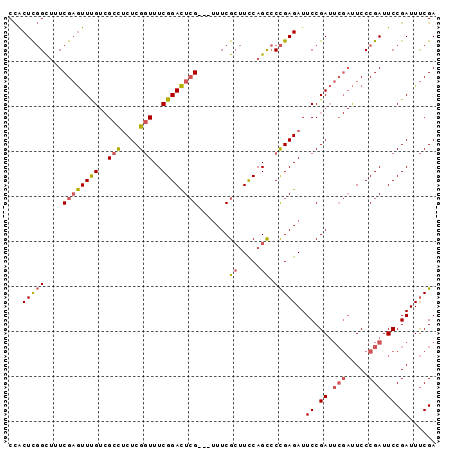
>2L_DroMel_CAF1 11359051 92 + 22407834 CCACUCGGCUUUCGAGUUUGUCGCUUCUCGGUUUCGGACUCG---UUUCGCUUCCAGCCCCGAGAUUCCGAUUCGAUUCCCGAUUCCGAUUUCGA ....((((...((((....((((..((((((..(.(((..((---...))..))).)..))))))...))))))))...))))...((....)). ( -24.80) >DroSec_CAF1 76261 92 + 1 CCACUCGGCUUUCGAGUUUGUCGCCUCUCGGUUUCGGACUCG---UUUCGCUUCCAGCCCCGAGAUUCCGAUUCGAUUCCCGAUUCCGAUUUCGA ......((((..((((((((..(((....)))..))))))))---..........)))).((((((.(.((.(((.....))).)).))))))). ( -26.30) >DroSim_CAF1 81016 92 + 1 CCACUCGGCUUUCGAGUUUGUCGCCUCUCGGUUUCGGACUCG---UUUCGCUUCCAGCCCCGAGAUUCCGAUUCGAUUCCCGAUUCCGAUUUCGA ......((((..((((((((..(((....)))..))))))))---..........)))).((((((.(.((.(((.....))).)).))))))). ( -26.30) >DroEre_CAF1 79813 92 + 1 CCACUCGGCUGUCGAGUUUGUCGCCUCUCGGUUUCGGACUCG---UUUCGCUUCCAGCCCCGAGAUUCCGAUUCGAUUCCCGAUUCCGAUUUCGA ......(((((.((((((((..(((....)))..))))))))---.........))))).((((((.(.((.(((.....))).)).))))))). ( -29.30) >DroAna_CAF1 68118 87 + 1 CUGCUUCGUAUUCGCAUUCG--GAUACUCGCUUUCGGAUUGGGAUAUUCGGAUUCAGU-CUGAGAUUCCGA-----CUCCCGAUUCCGAUUCCGA .......(((((((....))--)))))(((...(((((((((((...(((((.((...-....)).)))))-----.)))))).)))))...))) ( -31.30) >consensus CCACUCGGCUUUCGAGUUUGUCGCCUCUCGGUUUCGGACUCG___UUUCGCUUCCAGCCCCGAGAUUCCGAUUCGAUUCCCGAUUCCGAUUUCGA ...(((((....((((((((..(((....)))..)))))))).......((.....)).)))))..((.((.(((.....))).)).))...... (-17.68 = -18.24 + 0.56)

| Location | 11,359,051 – 11,359,143 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 84.76 |
| Mean single sequence MFE | -28.92 |
| Consensus MFE | -19.32 |
| Energy contribution | -21.16 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.776925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
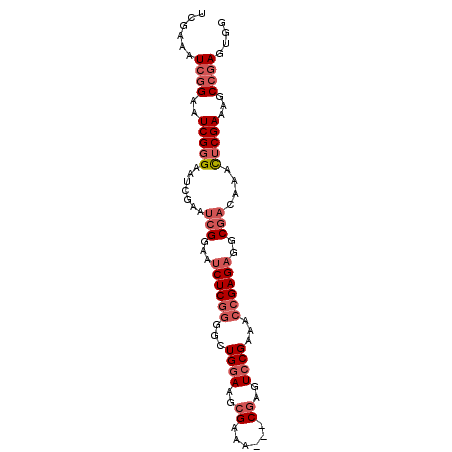
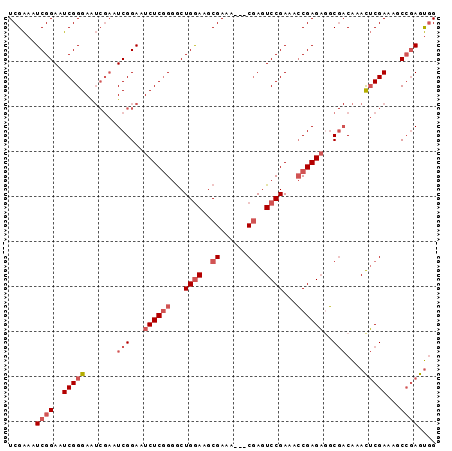
>2L_DroMel_CAF1 11359051 92 - 22407834 UCGAAAUCGGAAUCGGGAAUCGAAUCGGAAUCUCGGGGCUGGAAGCGAAA---CGAGUCCGAAACCGAGAAGCGACAAACUCGAAAGCCGAGUGG (((...((((...(((((.((......)).)))))(((((.(........---).)))))....))))....)))...(((((.....))))).. ( -28.50) >DroSec_CAF1 76261 92 - 1 UCGAAAUCGGAAUCGGGAAUCGAAUCGGAAUCUCGGGGCUGGAAGCGAAA---CGAGUCCGAAACCGAGAGGCGACAAACUCGAAAGCCGAGUGG .((...((((..(((((.......(((...((((((...((((..((...---))..))))...))))))..)))....)))))...)))).)). ( -28.90) >DroSim_CAF1 81016 92 - 1 UCGAAAUCGGAAUCGGGAAUCGAAUCGGAAUCUCGGGGCUGGAAGCGAAA---CGAGUCCGAAACCGAGAGGCGACAAACUCGAAAGCCGAGUGG .((...((((..(((((.......(((...((((((...((((..((...---))..))))...))))))..)))....)))))...)))).)). ( -28.90) >DroEre_CAF1 79813 92 - 1 UCGAAAUCGGAAUCGGGAAUCGAAUCGGAAUCUCGGGGCUGGAAGCGAAA---CGAGUCCGAAACCGAGAGGCGACAAACUCGACAGCCGAGUGG .((...((((..(((((.......(((...((((((...((((..((...---))..))))...))))))..)))....)))))...)))).)). ( -28.90) >DroAna_CAF1 68118 87 - 1 UCGGAAUCGGAAUCGGGAG-----UCGGAAUCUCAG-ACUGAAUCCGAAUAUCCCAAUCCGAAAGCGAGUAUC--CGAAUGCGAAUACGAAGCAG (((.(.(((((...((((.-----(((((.((....-...)).)))))...))))..(((....).))...))--))).).)))........... ( -29.40) >consensus UCGAAAUCGGAAUCGGGAAUCGAAUCGGAAUCUCGGGGCUGGAAGCGAAA___CGAGUCCGAAACCGAGAGGCGACAAACUCGAAAGCCGAGUGG ......((((..(((((.......(((...((((((...((((..((......))..))))...))))))..)))....)))))...)))).... (-19.32 = -21.16 + 1.84)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:33 2006