
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,349,551 – 11,349,650 |
| Length | 99 |
| Max. P | 0.992491 |

| Location | 11,349,551 – 11,349,650 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 89.03 |
| Mean single sequence MFE | -34.08 |
| Consensus MFE | -29.64 |
| Energy contribution | -29.20 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.17 |
| SVM RNA-class probability | 0.989491 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
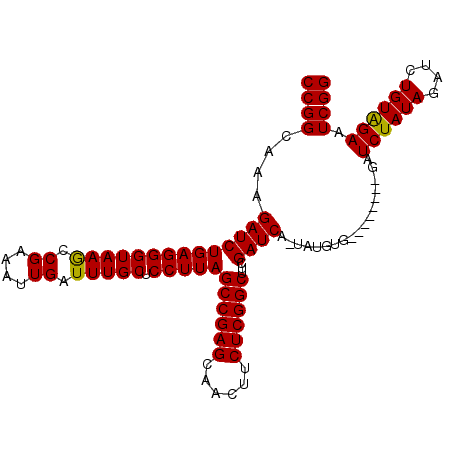
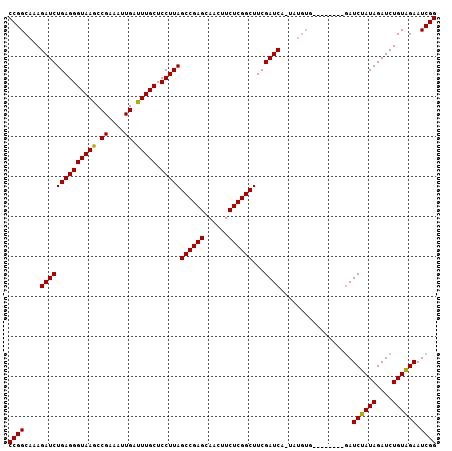
>2L_DroMel_CAF1 11349551 99 + 22407834 CCGGCAAAGAUCUGAGGGUAAGCCGAAAUUGAUUUGCUCCUUAGCCGAGCAACUUCUCGGCUUCGAUCAAUAUGUGG--------AUCUAUAGAUCUGUAGAAUCGG ((((....(((((((((((((..((....))..)))).)))))((((((......))))))...))))....((..(--------(((....))))..))...)))) ( -32.10) >DroSec_CAF1 66739 107 + 1 CCGGCAAAGAUCUGAGGGUAAGCCGAAAUUGAUUUGCUCCUUAGCCGAGCAACUUCUCGGCUUCGAUCAGUAUGUGGAUCUAUAGAUCUAUAGAUCUGUGGAAUCGG ((((....(((((((((((((..((....))..)))).)))))((((((......))))))...))))..........((((((((((....)))))))))).)))) ( -38.00) >DroSim_CAF1 71503 91 + 1 CCGGCAAAGAUCUGAGGGUAAGCCGAAAUUGAUUUGCUCCUUAGCCGAGCAACUUCUCGGCUUCGAUC----------------GAUCUAUAGAUCUGUGGAAUCGG ((((....(((((((((((((..((....))..)))).)))))((((((......))))))...))))----------------..((((((....)))))).)))) ( -29.50) >DroYak_CAF1 68208 99 + 1 CCGGCAAAGAUCUGAGGGUAAACCGAAAUUGAUUUGCUCCUUAGCCGAGCAACUUCUCGGCUUCGAUCGUUCUGUA--------GAUCUAUAGAUCUGUAGAAUCGG (((.....((((((((((((((.((....)).))))).)))))((((((......))))))...))))((((((((--------((((....))))))))))))))) ( -36.70) >consensus CCGGCAAAGAUCUGAGGGUAAGCCGAAAUUGAUUUGCUCCUUAGCCGAGCAACUUCUCGGCUUCGAUCA_UAUGUG________GAUCUAUAGAUCUGUAGAAUCGG ((((....((((((((((((((.((....)).))))).)))))((((((......))))))...))))..................((((((....)))))).)))) (-29.64 = -29.20 + -0.44)

| Location | 11,349,551 – 11,349,650 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 89.03 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -27.78 |
| Energy contribution | -27.59 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.33 |
| SVM RNA-class probability | 0.992491 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
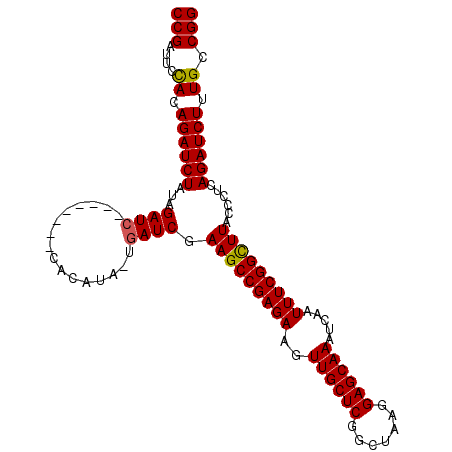
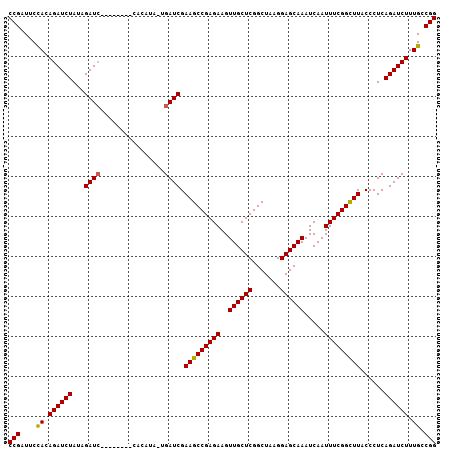
>2L_DroMel_CAF1 11349551 99 - 22407834 CCGAUUCUACAGAUCUAUAGAU--------CCACAUAUUGAUCGAAGCCGAGAAGUUGCUCGGCUAAGGAGCAAAUCAAUUUCGGCUUACCCUCAGAUCUUUGCCGG (((....((.((((((...(((--------(........)))).(((((((((..((((((.......)))))).....)))))))))......)))))).)).))) ( -29.60) >DroSec_CAF1 66739 107 - 1 CCGAUUCCACAGAUCUAUAGAUCUAUAGAUCCACAUACUGAUCGAAGCCGAGAAGUUGCUCGGCUAAGGAGCAAAUCAAUUUCGGCUUACCCUCAGAUCUUUGCCGG (((........((((((((....))))))))......((((...(((((((((..((((((.......)))))).....)))))))))....))))........))) ( -35.00) >DroSim_CAF1 71503 91 - 1 CCGAUUCCACAGAUCUAUAGAUC----------------GAUCGAAGCCGAGAAGUUGCUCGGCUAAGGAGCAAAUCAAUUUCGGCUUACCCUCAGAUCUUUGCCGG (((........((((....))))----------------((((.(((((((((..((((((.......)))))).....))))))))).......)))).....))) ( -29.00) >DroYak_CAF1 68208 99 - 1 CCGAUUCUACAGAUCUAUAGAUC--------UACAGAACGAUCGAAGCCGAGAAGUUGCUCGGCUAAGGAGCAAAUCAAUUUCGGUUUACCCUCAGAUCUUUGCCGG (((.((((..(((((....))))--------)..)))).((((..(((((((......))))))).(((.(.(((((......))))).))))..)))).....))) ( -30.80) >consensus CCGAUUCCACAGAUCUAUAGAUC________CACAUA_UGAUCGAAGCCGAGAAGUUGCUCGGCUAAGGAGCAAAUCAAUUUCGGCUUACCCUCAGAUCUUUGCCGG (((....((.((((((...((((................)))).(((((((((..((((((.......)))))).....)))))))))......)))))).)).))) (-27.78 = -27.59 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:30 2006