
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,304,662 – 11,304,762 |
| Length | 100 |
| Max. P | 0.740897 |

| Location | 11,304,662 – 11,304,762 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.23 |
| Mean single sequence MFE | -28.44 |
| Consensus MFE | -20.41 |
| Energy contribution | -20.27 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740897 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11304662 100 + 22407834 --------CA--CGUCAGUUGGACGCC-GUGUUUUGUCUAUCAUAAUGAGUGGCAGGCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCUGCCA---------GCUC --------..--.....(.((((((..-......)))))).).....((((((((((((((((.(((((((....)))))))......))))......)))))))))---------.))) ( -29.70) >DroVir_CAF1 23580 106 + 1 -------------AUCGGCCGGACGCC-GUCUUUUGUCUAUCAUAAUGAGUGGCAGGCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCUAGGACUGUGCCUAGUUC -------------..((((.....)))-)..............((((((((((((((.......(((((((....)))))))....)))))))))))))).(((((......)))))... ( -28.60) >DroGri_CAF1 28436 107 + 1 ------------CGUCAGUCGGACGCU-GUGUUUUGUCUAUCAUAAUGAGUGGCAGGCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCAGGGAUUGUGCGUAGUUU ------------.(((.....)))(((-((((...((((....((((((((((((((.......(((((((....)))))))....))))))))))))))....))))...))))))).. ( -29.40) >DroWil_CAF1 34242 107 + 1 CUAGCCUUGC--CGUCAUUUGGACGCC-GUGUUUUGUCUAUCAUAAUGAGUGGCAGUCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCAUUCU----------AUC ..(((..((.--((((.....)))).)-).)))..............((((((((((..((((.(((((((....)))))))......))))....)))))))))).----------... ( -26.30) >DroMoj_CAF1 33350 109 + 1 ---------C--CGUCGGACGGACGCCAGAGUUUUGUCUAUCAUAAUGAGUGGCAGGCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCGGGCACAGAGCUUAAUGC ---------.--((((.....))))...(((((((((((....((((((((((((((.......(((((((....)))))))....))))))))))))))...)).)))))))))..... ( -32.20) >DroPer_CAF1 35488 102 + 1 --------CCGUCGUCAGUCGGACGCC-GUGUUUUGUCCAUCAUAAUGAGUGGCAGUCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCUUGUCU---------UCAG --------..(..(.(((..(((((..-......)))))....(((((((((((((........(((((((....))))))).....))))))))))))).))).).---------.).. ( -24.42) >consensus _________C__CGUCAGUCGGACGCC_GUGUUUUGUCUAUCAUAAUGAGUGGCAGGCGGACAUUUGGACAUAAUUGUUCAAUUAUUUUGUUACUUAUUGCCUGGCA_________GUUC ....................(((((.........)))))....(((((((((((((........(((((((....))))))).....))))))))))))).................... (-20.41 = -20.27 + -0.14)

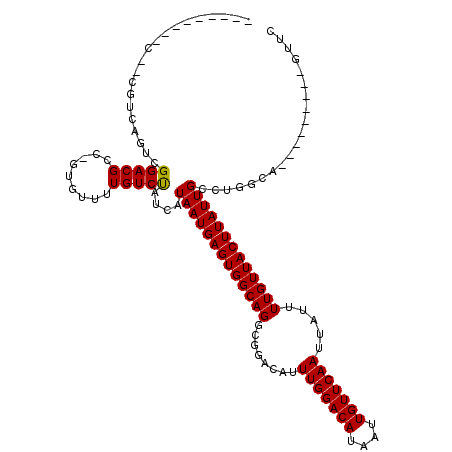
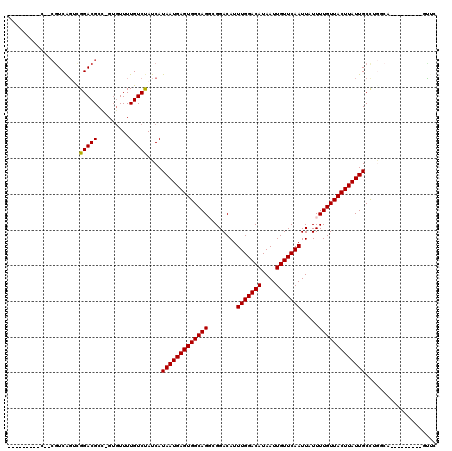
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:07 2006