| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,302,514 – 11,302,622 |
| Length | 108 |
| Max. P | 0.998602 |

| Location | 11,302,514 – 11,302,622 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 64.56 |
| Mean single sequence MFE | -24.41 |
| Consensus MFE | -12.85 |
| Energy contribution | -14.60 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.53 |
| SVM decision value | 3.16 |
| SVM RNA-class probability | 0.998602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11302514 108 + 22407834 UUAAGGUAAGUUUUA-GUGGUUUGUUCUAUCAGUAGCAGACUAGUGCACUCGGGAAAAUAAGCACUAUCACUUAAAACUUAUAGCUUUAAUGCAGGACCUUCGAAAUUU ..(((((((((((((-(((((((((((.....).)))))))((((((..............)))))).))).))))))))...((......))...)))))........ ( -26.34) >DroSec_CAF1 22142 89 + 1 UUAAGGUAAGUUUUA-GUGGUUUGUUCUAUCAGGAGCAGACUAGUGCACUCGUGAAAAUGAGAACUAUCACUUAAAACUUAAAAC-------------------AAAGU ....(.(((((((((-((((((((((((....)))))))))((((...(((((....))))).)))).))).)))))))))...)-------------------..... ( -28.60) >DroSim_CAF1 22110 89 + 1 UUAAGGUAAGUUUUA-GUGGUGGGUUCUAUCAGGAGCAGACUAGUGCACUCGGGAAAAUGGGAACUAUCACUUAAAACUUAAAAC-------------------AAAGU ....(.(((((((((-((((((((((((....)))))...........((((......))))..))))))).)))))))))...)-------------------..... ( -23.30) >DroYak_CAF1 22038 87 + 1 UUAAGGUAAGUUUUUACUGGUGUGUUCCC----GGGCAAAUGGG------------------AGGUAUCACUUAAAACUAAAAACUCUGCUGUCGCAACUGCCCAAAAU ....((((((((((....((((..(((((----(......))))------------------))....)))).))))))........(((....)))..))))...... ( -19.40) >consensus UUAAGGUAAGUUUUA_GUGGUGUGUUCUAUCAGGAGCAGACUAGUGCACUCGGGAAAAUGAGAACUAUCACUUAAAACUUAAAAC___________________AAAGU ......(((((((((.(((((((((((......)))))))).......((((......))))......))).)))))))))............................ (-12.85 = -14.60 + 1.75)

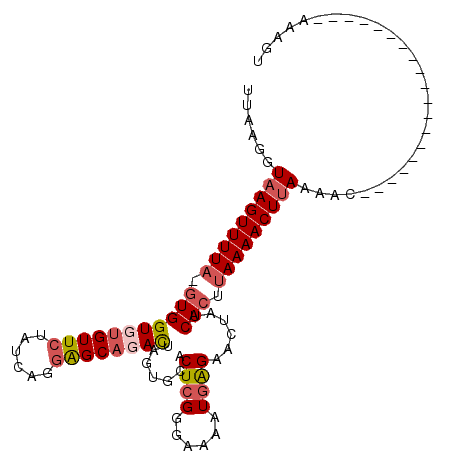
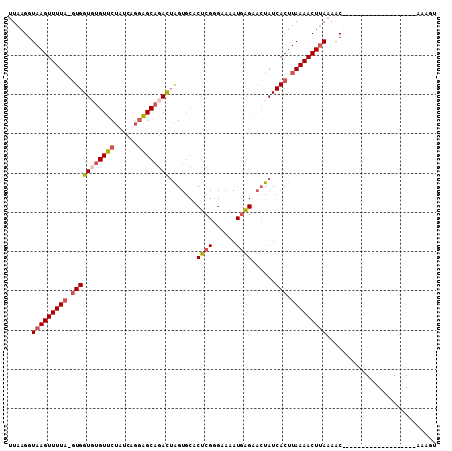
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:06 2006