
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,098,465 – 11,098,573 |
| Length | 108 |
| Max. P | 0.778278 |

| Location | 11,098,465 – 11,098,573 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 81.10 |
| Mean single sequence MFE | -41.16 |
| Consensus MFE | -22.73 |
| Energy contribution | -22.48 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778278 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11098465 108 - 22407834 CGAUGACGAUUCCAAUUCCAC---CAGUGCCCUUGUGCUGCACAUGUUUGUGGUGCAGGCAAAGAGCCUGCAGGAAUGUCAAAUAAUCUACCAGCCAUGUGUGGC------UGAAAA ...((((.(((((........---(((..(....)..)))((((....)))).(((((((.....))))))))))))))))..........((((((....))))------)).... ( -40.80) >DroVir_CAF1 2552 111 - 1 CGACGAUGAUAGCACUGCCACAACCGAUGCCCUAGUGGUGCAUAUGUUUGUGGUGCAGGCCAAGAGCUUGCAGGAGUGCCAGAUUAUCUAUCAGCCAUGUGUGGC------CGAAAA ....((((((.((((((((((((.((((((((....)).)))).)).)))))))((((((.....))))))...)))))...))))))..((.((((....))))------.))... ( -44.10) >DroPse_CAF1 2130 108 - 1 UGAUGAUGAGAACAAUUCCAC---AUACGCGCUGGUGGUGCACAUGUUUGUCGUGCAGGCCAAGAGCCUGCAGGAGUGCCACAUCAUCUACCAGCCGUGUGUGCC------CGAGAA ..................(((---(((((.((((((((.((((.......((.(((((((.....))))))).))))))........))))))))))))))))..------...... ( -42.91) >DroGri_CAF1 2061 111 - 1 UGAUGAUGAGAGCAAUGCCACCACCGAUGCGCUGGUGGUGCACAUGUUUGUCGUGCAGGCCAAGAGUCUGCAGGAGUGCCAGAUUAUCUAUCAGCCCUGUGUGCC------CGAAAA ..(((((..(((((.(((.(((((((......))))))))))..))))))))))((((((.....)))))).((.(..((((..............))).)..))------)..... ( -36.34) >DroAna_CAF1 1670 114 - 1 UGACGAGGACAACAACUCUAC---CUAUGCCCUGGUGUUGCACAUGUUCGUGGUGCAGGCCAAGAGCCUGCAAGAGUGCCAGAUAAUCUAUCAGCCAUGCAUGGCGGCAGCAGAAAA ....(((........)))...---.......(((.((((((.(((((..(((((((((((.....))))))..........((((...)))).)))))))))))))))).))).... ( -39.90) >DroPer_CAF1 2153 108 - 1 UGAUGAUGAGAACAAUUCCAC---AUACGCGCUGGUGGUGCACAUGUUUGUCGUGCAGGCCAAGAGCCUGCAGGAGUGCCACAUCAUCUACCAGCCGUGUGUGCC------CGAGAA ..................(((---(((((.((((((((.((((.......((.(((((((.....))))))).))))))........))))))))))))))))..------...... ( -42.91) >consensus UGAUGAUGAGAACAAUUCCAC___CUAUGCCCUGGUGGUGCACAUGUUUGUCGUGCAGGCCAAGAGCCUGCAGGAGUGCCAGAUAAUCUACCAGCCAUGUGUGCC______CGAAAA ....................................((((((((((.(((...(((((((.....)))))))(((...........)))..))).))))))))))............ (-22.73 = -22.48 + -0.25)

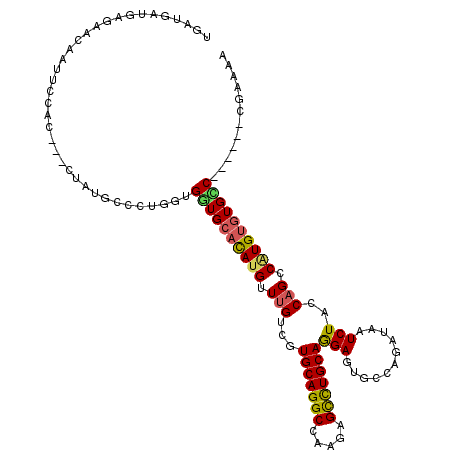

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:37 2006