
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,042,003 – 11,042,108 |
| Length | 105 |
| Max. P | 0.769888 |

| Location | 11,042,003 – 11,042,096 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 96.96 |
| Mean single sequence MFE | -21.10 |
| Consensus MFE | -17.34 |
| Energy contribution | -17.20 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.769888 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
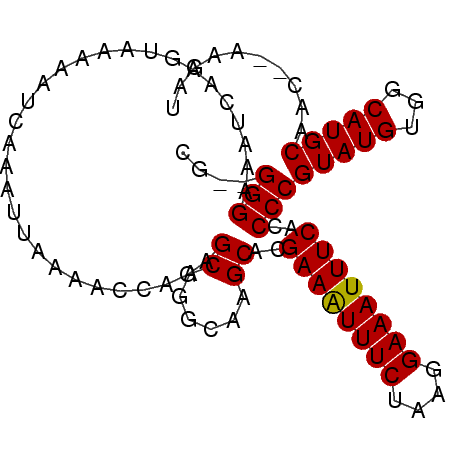
>2L_DroMel_CAF1 11042003 93 - 22407834 CG--GGGAAAUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC--AAGAU .(--(((..............................((......))..((((((((....)))))))).))))(((((...)))))...--..... ( -21.00) >DroPse_CAF1 48913 97 - 1 GGGGGGGAAAUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAACACAAGAU ..((((...............................((......))..((((((((....)))))))).))))...((((........)))).... ( -21.70) >DroSec_CAF1 30332 93 - 1 CG--GGGAAAUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAGUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC--AAGAU .(--(((..............................((......))..((((((((....)))))))).))))(((((...)))))...--..... ( -21.00) >DroEre_CAF1 32169 93 - 1 CG--GGGAAAUCAAGUAAAAAUCAAAUUAAAAUCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC--AAGAU .(--(((..............................((......))..((((((((....)))))))).))))(((((...)))))...--..... ( -21.00) >DroAna_CAF1 35999 92 - 1 CG--GGGA-AUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC--AAGAU .(--(((.-............................((......))..((((((((....)))))))).))))(((((...)))))...--..... ( -21.00) >DroPer_CAF1 48733 93 - 1 GG--GGGAAAUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC--AAGAU ((--((...............................((......))..((((((((....)))))))).))))(((((...)))))...--..... ( -20.90) >consensus CG__GGGAAAUCAAGUAAAAAUCAAAUUAAAACCAGAGCAGGCAAGCACGAAAUUUCUAAGGAAAUUUCACCCCGUAUGUGGCAUGCAAC__AAGAU ....(((..............................((......))..((((((((....))))))))..)))(((((...))))).......... (-17.34 = -17.20 + -0.14)


| Location | 11,042,018 – 11,042,108 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 92.61 |
| Mean single sequence MFE | -19.80 |
| Consensus MFE | -14.70 |
| Energy contribution | -15.45 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.668032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
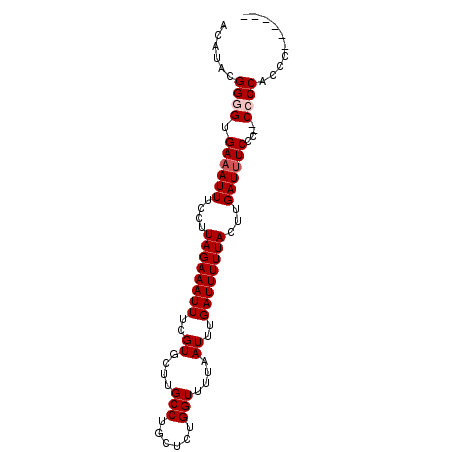
>2L_DroMel_CAF1 11042018 90 + 22407834 ACAUACGGGGUGAAAUUUCCUUAGAAAUUUCGUGCUUGCCUGCUCUGGUUUUAAUUUGAUUUUUACUUGAUUUCCC--CGCCACUCUACAUC .......(((.((((((....((((((((..((....(((......)))....))..))))))))...))))))))--)............. ( -18.40) >DroPse_CAF1 48930 86 + 1 ACAUACGGGGUGAAAUUUCCUUAGAAAUUUCGUGCUUGCCUGCUCUGGUUUUAAUUUGAUUUUUACUUGAUUUCCCCCCCCCGCCC------ ......((((.((((((....((((((((..((....(((......)))....))..))))))))...))))))))))........------ ( -21.10) >DroAna_CAF1 36014 83 + 1 ACAUACGGGGUGAAAUUUCCUUAGAAAUUUCGUGCUUGCCUGCUCUGGUUUUAAUUUGAUUUUUACUUGAU-UCCC--CGCCACUC------ .....(((((.((((((((....))))))))......(((......)))......................-.)))--))......------ ( -18.10) >DroPer_CAF1 48748 84 + 1 ACAUACGGGGUGAAAUUUCCUUAGAAAUUUCGUGCUUGCCUGCUCUGGUUUUAAUUUGAUUUUUACUUGAUUUCCC--CCCCGCCC------ .....(((((.((((((....((((((((..((....(((......)))....))..))))))))...))))))..--)))))...------ ( -21.60) >consensus ACAUACGGGGUGAAAUUUCCUUAGAAAUUUCGUGCUUGCCUGCUCUGGUUUUAAUUUGAUUUUUACUUGAUUUCCC__CCCCACCC______ ......((((.((((((....((((((((..((....(((......)))....))..))))))))...))))))....)))).......... (-14.70 = -15.45 + 0.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:01:00 2006