
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 11,025,542 – 11,025,636 |
| Length | 94 |
| Max. P | 0.986794 |

| Location | 11,025,542 – 11,025,636 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 85.60 |
| Mean single sequence MFE | -30.80 |
| Consensus MFE | -25.29 |
| Energy contribution | -25.13 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.854966 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11025542 94 + 22407834 UUCCAAG-ACCGGGGAACCCAACUCGAAAGUCAAGAUCCGGACUUGGGGACCCC------GGCCUGAGCCAAAGUAAAGAGUUGGUUGGGCGAUAAA-AAAU .......-..(((((..((((.......((((........))))))))..))))------)((((.((((((.........))))))))))......-.... ( -31.01) >DroSec_CAF1 14076 95 + 1 UUCCAAG-CCUGGGGAACCCAACUCGAAUGUCAAGAUCCGGACUUGGGGACCCC------GGCCUGAGCCAAAGUAACGAGUUGGUUGGGCAAUAAAAAAAU ...((.(-((.((((..(((((.(((.((......)).)))..)))))..))))------))).)).(((.((.(((....))).)).)))........... ( -29.80) >DroSim_CAF1 14345 93 + 1 UUCCAAG-CCCGGGGAACCCAACUCGAAAGUCA-GAUCCGGACUUGGGGACCCC------GGCCUGAGCCAAAGUAACGAGUUGGCUGGGCGAUAAA-AAAU ......(-(((((.....((((((((.(((((.-......)))))(((...)))------(((....))).......))))))))))))))......-.... ( -33.70) >DroEre_CAF1 15633 91 + 1 ----------UGGGGAACCCAGCUCGAGAGUCAAGAUCCGGACUUGGGGAGCCCGAGUUUGGCCUGGGCCAAAGUAACGAGUUGGCUGGGCGAUAAA-AAAU ----------.......(((((((((.((.......))))((((((((....)).(.((((((....)))))).)..))))))))))))).......-.... ( -30.60) >DroYak_CAF1 14873 95 + 1 UGGCAAAAGCAGGGGAGCCCAACUCGAAAGUCUAGAUCCGGACUUGGAGACCCC------GGCCUGAGCCAAAGUAACGAGUUGGCUGGGCGAUAAA-AAAU ..((...(((.((....))(((((((.((((((......))))))((.....))------(((....))).......))))))))))..))......-.... ( -28.90) >consensus UUCCAAG_CCCGGGGAACCCAACUCGAAAGUCAAGAUCCGGACUUGGGGACCCC______GGCCUGAGCCAAAGUAACGAGUUGGCUGGGCGAUAAA_AAAU ...........((((..(((((.......(((........))))))))..)))).......((((.((((((.........))))))))))........... (-25.29 = -25.13 + -0.16)


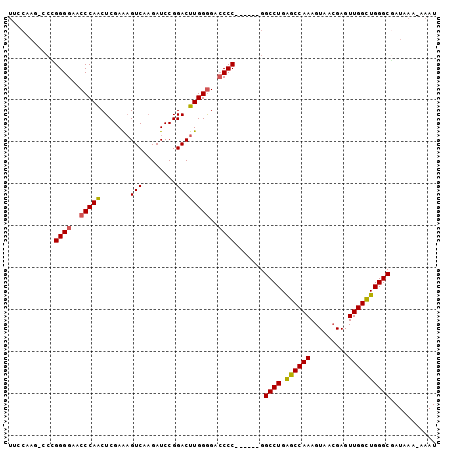
| Location | 11,025,542 – 11,025,636 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 85.60 |
| Mean single sequence MFE | -33.28 |
| Consensus MFE | -24.56 |
| Energy contribution | -25.40 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.74 |
| SVM decision value | 2.05 |
| SVM RNA-class probability | 0.986794 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 11025542 94 - 22407834 AUUU-UUUAUCGCCCAACCAACUCUUUACUUUGGCUCAGGCC------GGGGUCCCCAAGUCCGGAUCUUGACUUUCGAGUUGGGUUCCCCGGU-CUUGGAA ....-........(((.((((.........))))...(((((------((((..(((((.((.(((........))))).)))))..)))))))-))))).. ( -35.30) >DroSec_CAF1 14076 95 - 1 AUUUUUUUAUUGCCCAACCAACUCGUUACUUUGGCUCAGGCC------GGGGUCCCCAAGUCCGGAUCUUGACAUUCGAGUUGGGUUCCCCAGG-CUUGGAA ...........(((.((..((....))..)).)))(((((((------((((..(((((.((.((((......)))))).)))))..)))).))-))))).. ( -34.20) >DroSim_CAF1 14345 93 - 1 AUUU-UUUAUCGCCCAGCCAACUCGUUACUUUGGCUCAGGCC------GGGGUCCCCAAGUCCGGAUC-UGACUUUCGAGUUGGGUUCCCCGGG-CUUGGAA ....-........((((((((.........)))))..((.((------((((..(((((.((.(((..-.....))))).)))))..)))))).-))))).. ( -37.00) >DroEre_CAF1 15633 91 - 1 AUUU-UUUAUCGCCCAGCCAACUCGUUACUUUGGCCCAGGCCAAACUCGGGCUCCCCAAGUCCGGAUCUUGACUCUCGAGCUGGGUUCCCCA---------- ....-......(((((((......((((.((((((....)))))).(((((((.....)))))))....))))......)))))))......---------- ( -30.30) >DroYak_CAF1 14873 95 - 1 AUUU-UUUAUCGCCCAGCCAACUCGUUACUUUGGCUCAGGCC------GGGGUCUCCAAGUCCGGAUCUAGACUUUCGAGUUGGGCUCCCCUGCUUUUGCCA ....-..........................((((..((((.------((((..(((((.((.(((........))))).)))))..)))).))))..)))) ( -29.60) >consensus AUUU_UUUAUCGCCCAGCCAACUCGUUACUUUGGCUCAGGCC______GGGGUCCCCAAGUCCGGAUCUUGACUUUCGAGUUGGGUUCCCCAGG_CUUGGAA ...........(((..(((((.........)))))...))).......((((..(((((.((.(((........))))).)))))..))))........... (-24.56 = -25.40 + 0.84)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:52 2006