
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,982,694 – 10,982,814 |
| Length | 120 |
| Max. P | 0.724877 |

| Location | 10,982,694 – 10,982,814 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.82 |
| Mean single sequence MFE | -37.60 |
| Consensus MFE | -24.16 |
| Energy contribution | -24.38 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724877 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10982694 120 + 22407834 GGAAUAGAGUGCAAGUUCGUACCCACGAGGGUAGCUAGCUUCUUAGCGUACUGGCAAACUGCUGGCACUCGUGUGGUGCCCGACCAAUUGUAGUACAAGUGGCACAUCUUGUACGUAAGU ........(((((((...(((((((((((....((((((......((......)).....)))))).)))))).)))))..(.(((.(((.....))).))))....)))))))...... ( -41.20) >DroSec_CAF1 1 94 + 1 --------------------------GAGGGUAGCUAGCUUCUUAGCGUACUGGCAAACGGCGGGCACUCGUGUGGUGCCUGACCAAUUGUAGUACAAGUGGCACAUCUUGUACGUAAGU --------------------------((((((.(((((((....)))((((((.(((..((((((((((.....)))))))).))..)))))))))...)))))).))))..((....)) ( -33.40) >DroSim_CAF1 1 94 + 1 --------------------------GAGGGUAGCUAGCUUCUUAGCGUACUGGCAAACGGCGGGCACUCGUGUGGUGCCUGACCAAUUGUAGUACAAGUGGCACAUCUUGUACGUAAGU --------------------------((((((.(((((((....)))((((((.(((..((((((((((.....)))))))).))..)))))))))...)))))).))))..((....)) ( -33.40) >DroYak_CAF1 38 120 + 1 GGAAUGGAGUGCAAGUUCGUACCGACAAGGGUAGCAAGUUUUUUAGCGUACUGGCAAACGGCGGGCACCCGUGUGGUGCCUGACCAAUUAUAGUACAAGUGACACAUCUUGUACGUAAGC ........(((((((...((.((.....))((.((..(((....)))((((((......((((((((((.....)))))))).)).....))))))..)).))))..)))))))...... ( -37.10) >DroMoj_CAF1 41 120 + 1 GGUAUAGCAUGCAGAUGAGAACCAACAAGCGUAGCCAGUUUCUUAGCAUAUUGGCAAACAGCUGGCACCCGAGUGGUGCCCGACCAGUUGUAGUACAAAUGGCACAUUUUGUACGUUAGC (((....(((....)))...)))....(((...((((((..........))))))..((((((((((((.....)))))).....)))))).(((((((........))))))))))... ( -32.80) >DroAna_CAF1 41 120 + 1 GGAAUGGCGUGCAAGUUGGUGGCCACCAGGGUGGCCAGCUUCUUGGCAUACUGGCAGACGGCAGGCACCCGGGUAGUGCCCGACCAGUUGUAGUACAAGUGGCACAUUUUGUAAGUCAAC ....((((.((((((.((.(((((((....)))))))(((.((((...(((((.(((..(((.(((((.......))))).).))..)))))))))))).))).)).)))))).)))).. ( -47.70) >consensus GGAAU_G_GUGCAAGUU_GUACC_ACGAGGGUAGCUAGCUUCUUAGCGUACUGGCAAACGGCGGGCACCCGUGUGGUGCCCGACCAAUUGUAGUACAAGUGGCACAUCUUGUACGUAAGC .............................(((((((((....))))).)))).((((..(((.((((((.....)))))).).))..)))).(((((((........)))))))...... (-24.16 = -24.38 + 0.23)

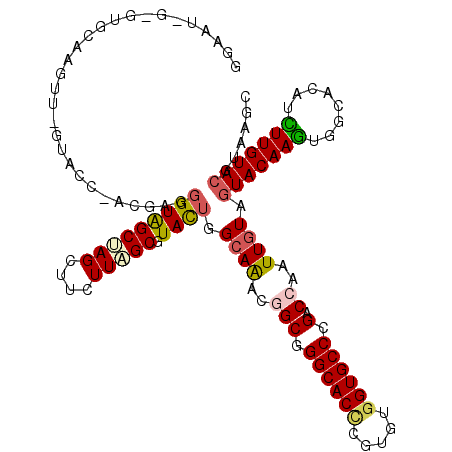
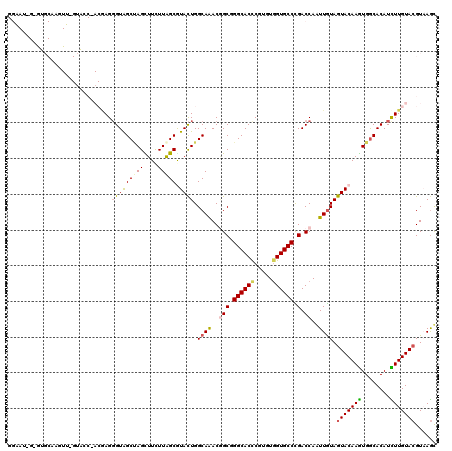
| Location | 10,982,694 – 10,982,814 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.82 |
| Mean single sequence MFE | -31.82 |
| Consensus MFE | -21.14 |
| Energy contribution | -21.20 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10982694 120 - 22407834 ACUUACGUACAAGAUGUGCCACUUGUACUACAAUUGGUCGGGCACCACACGAGUGCCAGCAGUUUGCCAGUACGCUAAGAAGCUAGCUACCCUCGUGGGUACGAACUUGCACUCUAUUCC ......(((((((........)))))))......((((.....))))...((((((....((((((.......((((......))))(((((....))))))))))).))))))...... ( -34.00) >DroSec_CAF1 1 94 - 1 ACUUACGUACAAGAUGUGCCACUUGUACUACAAUUGGUCAGGCACCACACGAGUGCCCGCCGUUUGCCAGUACGCUAAGAAGCUAGCUACCCUC-------------------------- ......((((.....)))).....(((((.(((.((((..((((((....).))))).)))).)))..)))))((((......)))).......-------------------------- ( -25.30) >DroSim_CAF1 1 94 - 1 ACUUACGUACAAGAUGUGCCACUUGUACUACAAUUGGUCAGGCACCACACGAGUGCCCGCCGUUUGCCAGUACGCUAAGAAGCUAGCUACCCUC-------------------------- ......((((.....)))).....(((((.(((.((((..((((((....).))))).)))).)))..)))))((((......)))).......-------------------------- ( -25.30) >DroYak_CAF1 38 120 - 1 GCUUACGUACAAGAUGUGUCACUUGUACUAUAAUUGGUCAGGCACCACACGGGUGCCCGCCGUUUGCCAGUACGCUAAAAAACUUGCUACCCUUGUCGGUACGAACUUGCACUCCAUUCC ((.((((((((((........)))))))..(((.((((..((((((.....)))))).)))).)))...))).))........(((.((((......)))))))................ ( -30.70) >DroMoj_CAF1 41 120 - 1 GCUAACGUACAAAAUGUGCCAUUUGUACUACAACUGGUCGGGCACCACUCGGGUGCCAGCUGUUUGCCAAUAUGCUAAGAAACUGGCUACGCUUGUUGGUUCUCAUCUGCAUGCUAUACC ((.((((((((((........))))))).......(((..((((((.....)))))).)))))).))...(((((...(((((..((.......))..))).))....)))))....... ( -32.40) >DroAna_CAF1 41 120 - 1 GUUGACUUACAAAAUGUGCCACUUGUACUACAACUGGUCGGGCACUACCCGGGUGCCUGCCGUCUGCCAGUAUGCCAAGAAGCUGGCCACCCUGGUGGCCACCAACUUGCACGCCAUUCC ((.(((.........((((.....)))).......((.((((((((.....))))))))))))).))..(..(((.(((..(.(((((((....))))))).)..))))))..)...... ( -43.20) >consensus ACUUACGUACAAGAUGUGCCACUUGUACUACAAUUGGUCAGGCACCACACGAGUGCCCGCCGUUUGCCAGUACGCUAAGAAGCUAGCUACCCUCGU_GGUAC_AACUUGCAC_C_AUUCC ......(((((((........)))))))....(((((((.(((((.......))))).)......))))))..((((......))))................................. (-21.14 = -21.20 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:44 2006