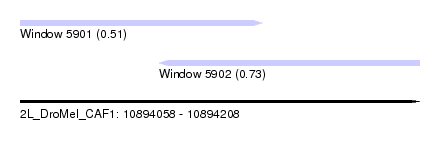
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,894,058 – 10,894,208 |
| Length | 150 |
| Max. P | 0.727169 |

| Location | 10,894,058 – 10,894,149 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 89.07 |
| Mean single sequence MFE | -22.25 |
| Consensus MFE | -18.91 |
| Energy contribution | -19.35 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.505785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
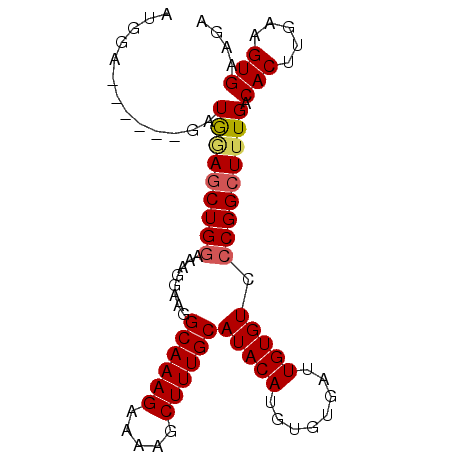
>2L_DroMel_CAF1 10894058 91 + 22407834 CUGGAGCUGUAGAUGGAACUGAAAGGGAAGGCAAAGAAAAGCUUUGCAUACAUGUGUGAUUGUGUCCCGGCUUUGACACUUGAAGUGAAGA ..((((((.(((......)))...((((..((((((.....)))))).((((........))))))))))))))..(((.....))).... ( -21.80) >DroSec_CAF1 33622 85 + 1 AUGGA------GCUAUAGCUGGAAAGGAAGGCAAAGAAAAGCUUUGCAUACAUGUGUGAUUGUGUCCCGGCUUUGACACUUGAAGUGAAGA .....------..((.((((((........((((((.....))))))(((((........))))).)))))).)).(((.....))).... ( -19.90) >DroSim_CAF1 34505 91 + 1 AUGGAGCUGAAGCUGAAGCUGGAAAGGAAGGCAAAGAAAAGCUUUGCAUACAUGUGUGAUUGUGUCCCGGCUUUGACACUUGAAGUGAAGA ....(((....)))((((((((........((((((.....))))))(((((........))))).))))))))..(((.....))).... ( -24.80) >DroEre_CAF1 33827 85 + 1 CCGGA------GAUGGAGCUGGAGAGGAAGGCAAAGAAAAGCUUUGCAUACAUGUGUGAUUGUGUCCCGGCUUUGACACUUGAAGUGGAGA .....------...((((((((.((.....((((((.....)))))).((((........))))))))))))))..(((.....))).... ( -22.50) >consensus AUGGA______GAUGGAGCUGGAAAGGAAGGCAAAGAAAAGCUUUGCAUACAUGUGUGAUUGUGUCCCGGCUUUGACACUUGAAGUGAAGA .............(((((((((........((((((.....))))))(((((........))))).))))))))).(((.....))).... (-18.91 = -19.35 + 0.44)

| Location | 10,894,110 – 10,894,208 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 89.80 |
| Mean single sequence MFE | -25.32 |
| Consensus MFE | -20.44 |
| Energy contribution | -20.48 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.727169 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
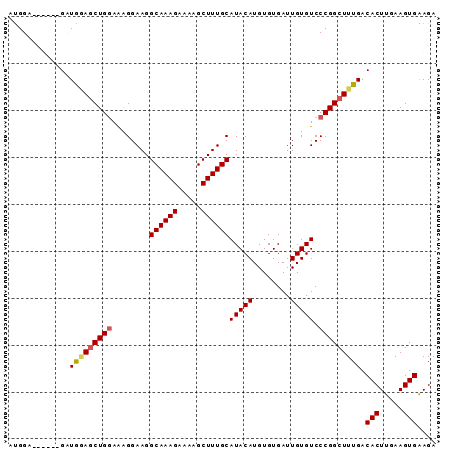
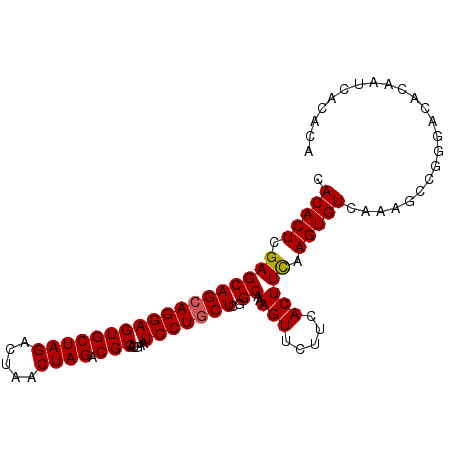
>2L_DroMel_CAF1 10894110 98 - 22407834 AACACUCGAGCAGCAGGAGUGCUAGACUAACUAGACGCAUGAAUCCUACUCGGCAAAGUUCUUCACUUCAAGUGUCAAAGCCGGGACACAAUCACACA ....((((.((((.(((((((((((.....)))).))).....)))).)).((((((((.....))))....))))...))))))............. ( -19.70) >DroSec_CAF1 33668 98 - 1 CACACUCGAGCAGCAGGAGUGCUAGACUAACUAGACGCAUGAAUCCUGCUCGGCAAAGUUCUUCACUUCAAGUGUCAAAGCCGGGACACAAUCACACA ....((((.((((((((((((((((.....)))).))).....))))))).((((((((.....))))....))))...))))))............. ( -26.30) >DroSim_CAF1 34557 98 - 1 CACACUCGAGCAGCAGGAGUGCUAGGCUAACUAGACGCAUGAAUCCUGCUCGGCAAAGUUCUUCACUUCAAGUGUCAAAGCCGGGACACAAUCACACA ....((((.((((((((((((((((.....)))).))).....))))))).((((((((.....))))....))))...))))))............. ( -27.60) >DroEre_CAF1 33873 98 - 1 CACACUCGAGCAGCAGGAGUGCUAGACUAACUAGACGCAUGAAUCCUGCUCGGCAGAGUUCUCCACUUCAAGUGUCAAAGCCGGGACACAAUCACACA ...((((..((((((((((((((((.....)))).))).....)))))))..)).))))............(((((........)))))......... ( -27.70) >DroYak_CAF1 34660 81 - 1 CACACUCGAGCAGCAGGAGUGCUAGACUAACUAGACGCAUGAAUCCUGCUCGGCCGAGUUCUCCACUUUAAGUGUC-----------------ACACA ...(((((.((((((((((((((((.....)))).))).....)))))))..)))))))....(((.....)))..-----------------..... ( -25.30) >consensus CACACUCGAGCAGCAGGAGUGCUAGACUAACUAGACGCAUGAAUCCUGCUCGGCAAAGUUCUUCACUUCAAGUGUCAAAGCCGGGACACAAUCACACA .(((((.((((((((((((((((((.....)))).))).....)))))))..))..(((.....))))).)))))....................... (-20.44 = -20.48 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:00:04 2006