
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,869,467 – 10,869,573 |
| Length | 106 |
| Max. P | 0.886638 |

| Location | 10,869,467 – 10,869,573 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 71.90 |
| Mean single sequence MFE | -18.26 |
| Consensus MFE | -6.93 |
| Energy contribution | -7.93 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.38 |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.886638 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
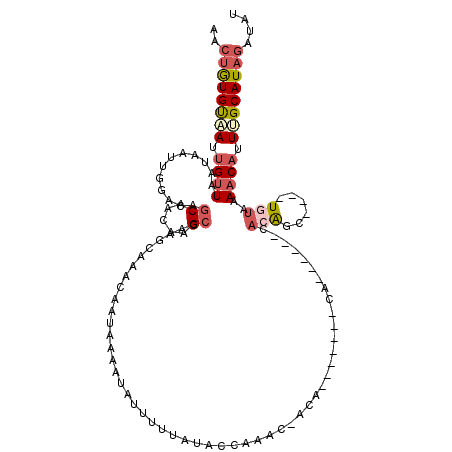
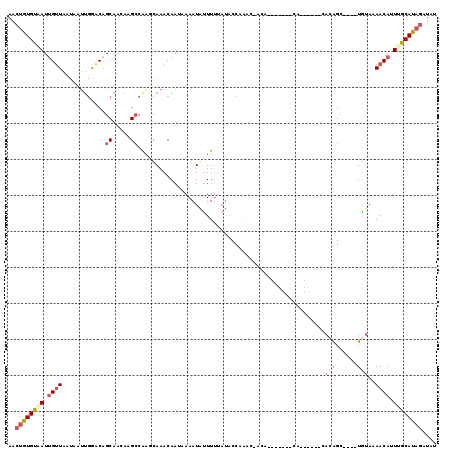
>2L_DroMel_CAF1 10869467 106 - 22407834 AACUGUGUAAUUGUUAAUAAUUGGACAGCAAUACGGCAAGUAAACAAAUAAAUAUUUUUAUACCCAACCACA-------CA------UACAGCCGUAUAAAAAACAUUUGCAUAGAUAU ..((((((((.((((......((.....))(((((((..(....)..(((((....)))))...........-------..------....)))))))....)))).)))))))).... ( -20.80) >DroSec_CAF1 16234 101 - 1 AACUGUGUUAUUGUUAAUAAUUGGACAGCAACACGCCAAGUAAACAAUAAAAUAUUUUUAUACCCAAC-ACA-------CA------CACGGC----UAUAAAACAUUUGCAUAGAUAU ..(((((((((((((.....((((.(........)))))...)))))))......(((((((.((...-...-------..------...)).----))))))).....)))))).... ( -16.80) >DroSim_CAF1 16632 107 - 1 AACUGUGUAAUUGUUAAUAAUUGGACAGCAAAACGCCAAGUAAACAAUAAAAUAUUUUUAUACCCAAC-ACACACACA-CA------CACGGC----UGUAAAACAUUUGCAUAGAUAU ..(((((((((((((.....((((.(........)))))...)))))).......(((((((.((...-.........-..------...)).----)))))))....))))))).... ( -16.73) >DroEre_CAF1 16618 89 - 1 AACUGUGCGAUGGCUAC------AACUGCAACAAGCCAAUCAAACAAUGAAAUGCUUUAUGCCCA--------------CA------CACAGC----UGUAAAACAUUCGCAUAGAUGU ..((((((((((..(((------(.(((((..((((...(((.....)))...))))..)))...--------------..------...)).----))))...)).)))))))).... ( -20.30) >DroYak_CAF1 16670 95 - 1 AACUGUGUAAUUGUUAUUAAUUGGACUGCAUCAAGCCAAGCAAACAAUGAAAUAUUUGUAUACCA--------------AA------CACAGC----UGUAAAACAUUCGCAUAGAUAU ..(((((..(.((((.....((((.((......))))))....((((........))))......--------------..------......----.....)))).)..))))).... ( -15.40) >DroAna_CAF1 17690 113 - 1 AAGCAUGUAAUUGUUAAUAAUUGAGCAGCAACAAGCCGGACAAACAA-AAAAUAUUUUCAUUCCAAAA-UUGAAUAAAGCAUUUGGUGACAGA----UGUACAACAUUUGCAUAGAUAU ....((((((.((((..((((((.((........)))(((.......-.............)))..))-)))......(((((((....))))----)))..)))).))))))...... ( -19.55) >consensus AACUGUGUAAUUGUUAAUAAUUGGACAGCAACAAGCCAAGCAAACAAUAAAAUAUUUUUAUACCAAAC_ACA_______CA______CACAGC____UGUAAAACAUUUGCAUAGAUAU ..((((((((.((((............((.....))....................................................(((......)))..)))).)))))))).... ( -6.93 = -7.93 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:43 2006