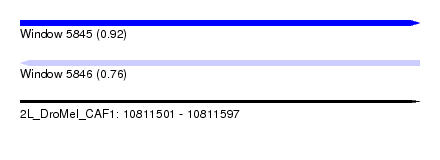

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,811,501 – 10,811,597 |
| Length | 96 |
| Max. P | 0.919616 |

| Location | 10,811,501 – 10,811,597 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 89.44 |
| Mean single sequence MFE | -37.68 |
| Consensus MFE | -32.72 |
| Energy contribution | -33.45 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10811501 96 + 22407834 GGCGUAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCUGCGCGCAUCGGUAUGCGUGCGAGG ......(((((......(((((....)))))......(((.(((((((((....)))))))))))).)))))((((((((....)))))))).... ( -43.30) >DroSec_CAF1 45214 96 + 1 GGCAGAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCUGCGCGCAUCGGUAUGUGUGCGAGG ......(((((......(((((....)))))......(((.(((((((((....)))))))))))).)))))((((((((....)))))))).... ( -41.40) >DroSim_CAF1 50609 96 + 1 GGCGGAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCUGUGCGCAUCGGUAUGUGUGCGAGG ......(((((......(((((....)))))......(((.(((((((((....)))))))))))).)))))(..(((((....)))))..).... ( -39.00) >DroEre_CAF1 46035 94 + 1 GGCGGAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCA--GUGCAUCGGUGUGUGUGCGAGU .......((((......(((((....)))))......(((.(((((((((....)))))))))))).)))).--(..(((......)))..).... ( -35.10) >DroYak_CAF1 47085 96 + 1 GGCGGAAGGGAAAUUGCGCGGAUUUGUCUGUUCUCUCGAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCUGCGUGCAUCGGUAUGUGUGCCAGU ((((..(((((....(.(((((....))))).)....(((.(((((((((....)))))))))))).)))))((((((....)))))).))))... ( -39.70) >DroAna_CAF1 45634 88 + 1 GGCAAAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGGGUCCUUAUCCUUUU--------CUAUGUGUCAGUGU (((((((((((......(((((....)))))......(((.(((.(((((....))))).)))))).)))))))--------.....))))..... ( -27.60) >consensus GGCGGAAGGGAAAUUGCGCGGAUUUGUCUGCUCUCUCAAGCGGCACGCACUUAAGUGCGUGUCCUUAUCCCUGCGCGCAUCGGUAUGUGUGCGAGG ......(((((......(((((....)))))......(((.(((((((((....)))))))))))).)))))((((((((....)))))))).... (-32.72 = -33.45 + 0.73)

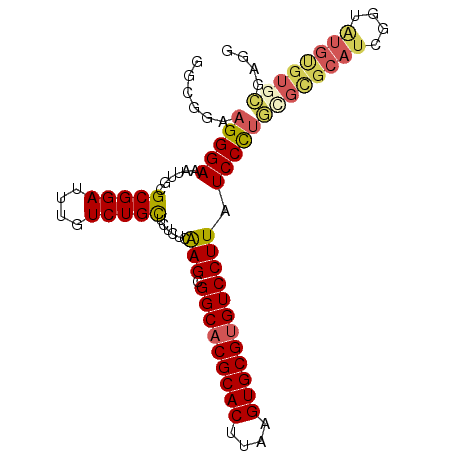
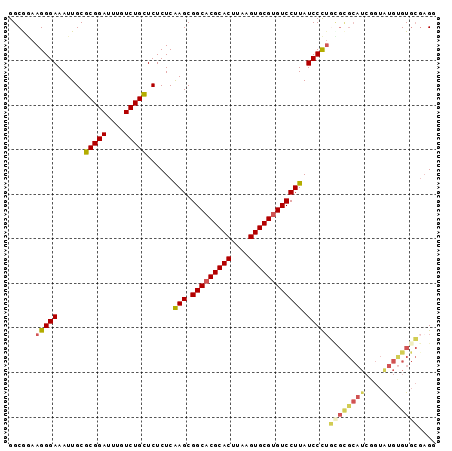
| Location | 10,811,501 – 10,811,597 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 89.44 |
| Mean single sequence MFE | -31.50 |
| Consensus MFE | -26.03 |
| Energy contribution | -26.00 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.759499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10811501 96 - 22407834 CCUCGCACGCAUACCGAUGCGCGCAGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUACGCC ....((..((((....))))((((..((((..((.(((((((....)))))))))((((....))))......))))))))............)). ( -38.40) >DroSec_CAF1 45214 96 - 1 CCUCGCACACAUACCGAUGCGCGCAGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUCUGCC ....((((.......).)))((((..((((..((.(((((((....)))))))))((((....))))......))))))))............... ( -34.40) >DroSim_CAF1 50609 96 - 1 CCUCGCACACAUACCGAUGCGCACAGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUCCGCC .................(((((....((((..((.(((((((....)))))))))((((....))))......))))))))).............. ( -32.80) >DroEre_CAF1 46035 94 - 1 ACUCGCACACACACCGAUGCAC--UGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUCCGCC ....((((.......).)))..--..((((..((.(((((((....)))))))))((((....))))......))))(((.((......)).))). ( -29.30) >DroYak_CAF1 47085 96 - 1 ACUGGCACACAUACCGAUGCACGCAGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUCGAGAGAACAGACAAAUCCGCGCAAUUUCCCUUCCGCC ...(((...(((....)))...(.(((((...((.(((((((....)))))))))((.((.((...........)))).))....))))).).))) ( -29.50) >DroAna_CAF1 45634 88 - 1 ACACUGACACAUAG--------AAAAGGAUAAGGACCCGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUUUGCC ..............--------....((((..((.(.(((((....))))).)))((((....))))......))))..((((........)))). ( -24.60) >consensus ACUCGCACACAUACCGAUGCGCGCAGGGAUAAGGACACGCACUUAAGUGCGUGCCGCUUGAGAGAGCAGACAAAUCCGCGCAAUUUCCCUUCCGCC ..........................((((..((.(((((((....)))))))))((((....))))......))))(((.((......)).))). (-26.03 = -26.00 + -0.03)

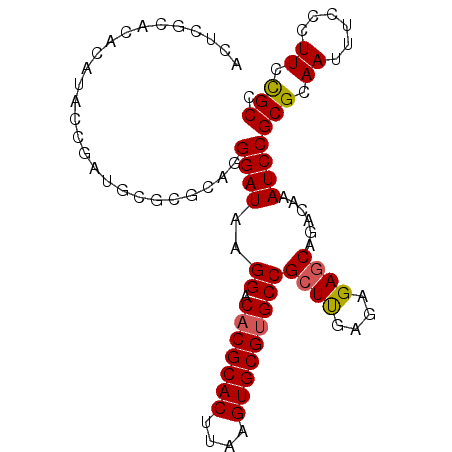
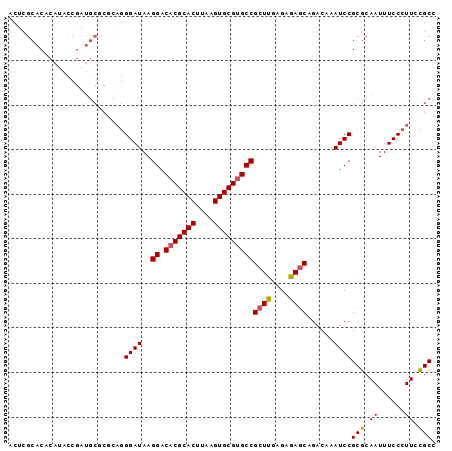
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:59:09 2006