| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,791,614 – 10,791,706 |
| Length | 92 |
| Max. P | 0.998424 |

| Location | 10,791,614 – 10,791,706 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 77.10 |
| Mean single sequence MFE | -37.73 |
| Consensus MFE | -24.06 |
| Energy contribution | -24.25 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.64 |
| SVM decision value | 3.10 |
| SVM RNA-class probability | 0.998424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10791614 92 - 22407834 GUCAGAUGGGAGAAG---AAGGAGCAGGAGGAGCUAG------AGGGGCCUACUCCUCUGGCGCACUCAGUUAGUCCAAAGCCCUUUCCUUUCUUUGGCUU ((((((.(..((..(---((((.((..(((..(((((------(((((....))))))))))...))).(......)...))))))).))..))))))).. ( -36.90) >DroSec_CAF1 25402 85 - 1 GCCAGAUGGGAGGAGG---------------AGCUGGAGCU-UAGGGGCUUCCUUCUCUGGCGGACUCAGUUAGUCCAAAGCCCUUUCCUUUCUUUGGCUU ((((((.(..((((((---------------(((....)))-).(((((((((......)).(((((.....))))).))))))))))))..))))))).. ( -38.60) >DroSim_CAF1 27024 101 - 1 GCCAGAUGGGAGGAGGAGCAGGAGCAGGAGGAGCUGGAGCUGUAGGGGCUUCCUUCUCUGGCGGACUCAGUUAGUCCAAAGCCCUUUCCUUUCUUUGGCUU ((((((.(..((((((.((....((.(((((((..((((((.....))))))))))))).))(((((.....)))))...))))..))))..))))))).. ( -46.10) >DroYak_CAF1 26383 86 - 1 GUCAGAUGGGCGGAGAAGAAGG---------------AGCAGGAGGGGCAUUCUCUUCUGGCGGACUCAGUUAGUCCAAAGCCCUUUCCUUUCUUUGGCUU .......((.((((((((((((---------------.(((((((((.....)))))))...(((((.....)))))...)))))))...))))))).)). ( -29.30) >consensus GCCAGAUGGGAGGAGG__AAGG_________AGCUGGAGCU_UAGGGGCUUCCUCCUCUGGCGGACUCAGUUAGUCCAAAGCCCUUUCCUUUCUUUGGCUU ((((((.(((((((((...........................(((((......)))))((((((((.....)))))...))).))))))))))))))).. (-24.06 = -24.25 + 0.19)

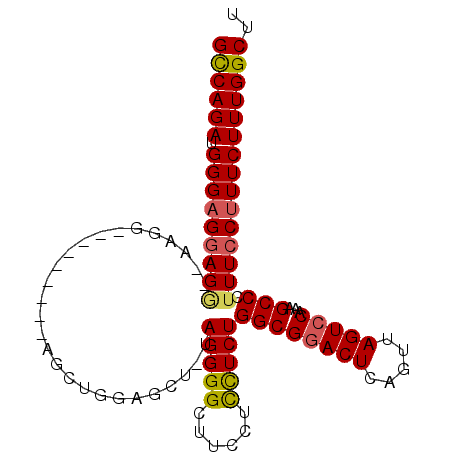
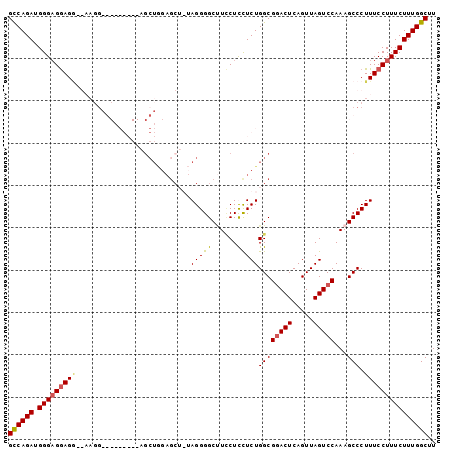
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:59 2006