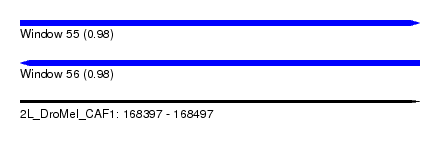

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 168,397 – 168,497 |
| Length | 100 |
| Max. P | 0.979340 |

| Location | 168,397 – 168,497 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 91.96 |
| Mean single sequence MFE | -18.98 |
| Consensus MFE | -16.34 |
| Energy contribution | -16.98 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979340 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
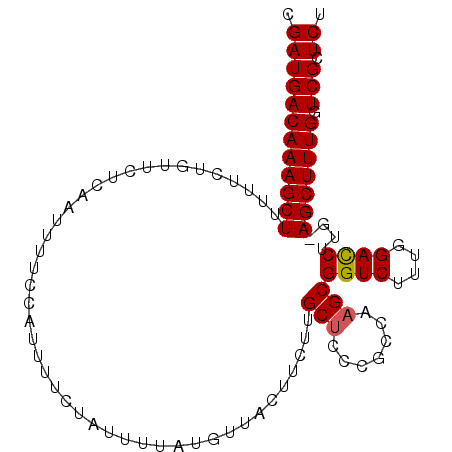
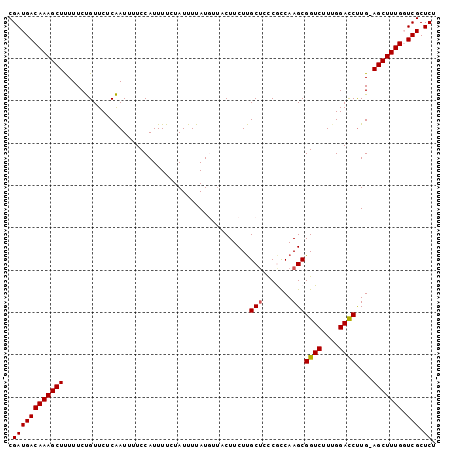
>2L_DroMel_CAF1 168397 100 + 22407834 CGAUGACAAAGCUUUUUCUGUUCUCAAUAUUCCAUUUUCUAUUUUAUGUUACUUCUUGCUCCCGCCAAGCGGUCUUCGGACCUUG-AGCUUUGUUCGCUCU .(((((((((((((...........................................(((.......)))((((....))))..)-))))))).))).)). ( -21.80) >DroSec_CAF1 14837 101 + 1 CGAUGACAAAGCUUUUUCUGUUCUCAAUUUUCCAUUUUCUAUUUUAUGUUACUUCUUGCUCCCGCCAAGCGGUCUUUGGAUCUUGUAGCUUUGCUCGCUCU .((((((((((((...........(((...((((.............((........))..((((...))))....))))..))).))))))).))).)). ( -16.90) >DroSim_CAF1 15001 101 + 1 CGAUGACAAAGCUUUUUCUGUUCUCAAUUUUCCAUUUUCUAUUUUAUGUUACUUCUUGCUCCCGCCAAGCGGUCUUUGGACCCUGUAGCUUUGGUCGCUCU .((((((((((((............................................(((.......)))((((....))))....)))))).)))).)). ( -17.10) >DroEre_CAF1 15042 100 + 1 CGAUGACAAAGCUCUUUCUGUUCUCAAUUUUCCAUUUUCUAUUUUAUUUCCCUUCUUGCACUCAUCAAGCGGUCUUUGGACCUUG-AGCUUUGGUCGCUCU .(((((((((((((........................................((((.......)))).((((....))))..)-)))))).)))).)). ( -20.00) >DroYak_CAF1 14892 100 + 1 CGAUGACAAAGCUUUUUCCGUUCUCGAUUUUCCAUUUUCUAUUUUAUUUUACUUCUCGCUCUCAUCAAGCGGUCUUUGGACCUUG-AGCUUUGGUCGCUCU .(((((((((((((...........................................(((.......)))((((....))))..)-)))))).)))).)). ( -19.10) >consensus CGAUGACAAAGCUUUUUCUGUUCUCAAUUUUCCAUUUUCUAUUUUAUGUUACUUCUUGCUCCCGCCAAGCGGUCUUUGGACCUUG_AGCUUUGGUCGCUCU .((((((((((((............................................(((.......)))((((....))))....))))))).))).)). (-16.34 = -16.98 + 0.64)

| Location | 168,397 – 168,497 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 91.96 |
| Mean single sequence MFE | -20.47 |
| Consensus MFE | -17.96 |
| Energy contribution | -17.76 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978758 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
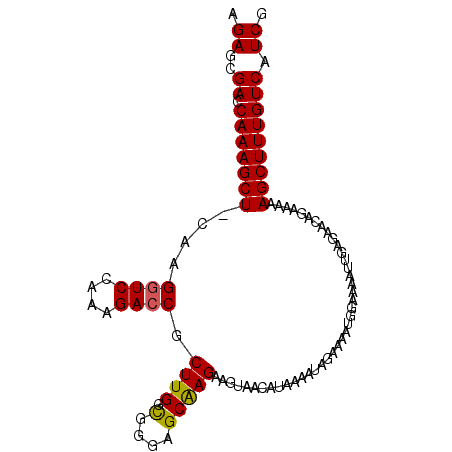
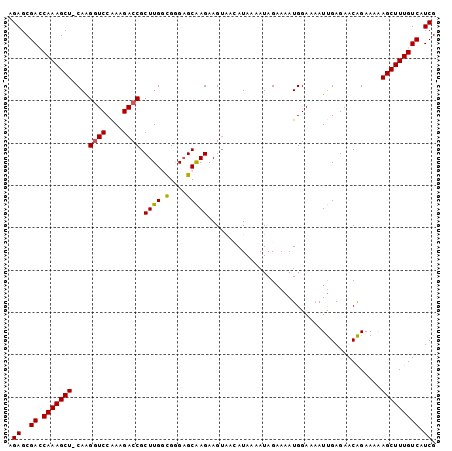
>2L_DroMel_CAF1 168397 100 - 22407834 AGAGCGAACAAAGCU-CAAGGUCCGAAGACCGCUUGGCGGGAGCAAGAAGUAACAUAAAAUAGAAAAUGGAAUAUUGAGAACAGAAAAAGCUUUGUCAUCG .((..((.(((((((-...((((....))))((((.((....))...)))).....................................))))))))).)). ( -20.90) >DroSec_CAF1 14837 101 - 1 AGAGCGAGCAAAGCUACAAGAUCCAAAGACCGCUUGGCGGGAGCAAGAAGUAACAUAAAAUAGAAAAUGGAAAAUUGAGAACAGAAAAAGCUUUGUCAUCG .((..((.(((((((......((((....((((...))))..((.....))................))))...(((....)))....))))))))).)). ( -18.40) >DroSim_CAF1 15001 101 - 1 AGAGCGACCAAAGCUACAGGGUCCAAAGACCGCUUGGCGGGAGCAAGAAGUAACAUAAAAUAGAAAAUGGAAAAUUGAGAACAGAAAAAGCUUUGUCAUCG .((..(((.((((((....((((....))))((((.((....))...)))).....................................))))))))).)). ( -21.30) >DroEre_CAF1 15042 100 - 1 AGAGCGACCAAAGCU-CAAGGUCCAAAGACCGCUUGAUGAGUGCAAGAAGGGAAAUAAAAUAGAAAAUGGAAAAUUGAGAACAGAAAGAGCUUUGUCAUCG .((..(((.((((((-(..((((....)))).((..((...(.((......................)).)..))..))........)))))))))).)). ( -22.05) >DroYak_CAF1 14892 100 - 1 AGAGCGACCAAAGCU-CAAGGUCCAAAGACCGCUUGAUGAGAGCGAGAAGUAAAAUAAAAUAGAAAAUGGAAAAUCGAGAACGGAAAAAGCUUUGUCAUCG .((..(((.((((((-...((((....))))((((.....))))............................................))))))))).)). ( -19.70) >consensus AGAGCGACCAAAGCU_CAAGGUCCAAAGACCGCUUGGCGGGAGCAAGAAGUAACAUAAAAUAGAAAAUGGAAAAUUGAGAACAGAAAAAGCUUUGUCAUCG .((..((.(((((((....((((....)))).((((.(....))))).........................................))))))))).)). (-17.96 = -17.76 + -0.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:23:53 2006