
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,354,425 – 10,354,543 |
| Length | 118 |
| Max. P | 0.751985 |

| Location | 10,354,425 – 10,354,543 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.49 |
| Mean single sequence MFE | -36.79 |
| Consensus MFE | -23.06 |
| Energy contribution | -24.88 |
| Covariance contribution | 1.81 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.677348 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10354425 118 + 22407834 CUGUUUUAUCGAUUUGUGGGCGGGCUCAGUGCCAUAAAAC--AAGGAUCUGAGUUGGCGCUUUGCCACGCCCCCCAUAUUCAUUACUUCAACUUUGUGCUAAUGACACUGGGCUACAAAG ............(((((((((((((..(((((((......--............)))))))..))).)))))((((...((((((...((....))...))))))...))))..))))). ( -38.07) >DroSim_CAF1 7125 118 + 1 CUGUUUUAUCGAUUUGUGGGCGGGCUCAGUGCCAUAAAAC--AUGGAUCUGAGUUGGCGCUUUGCCACGCCCCCCAUAUUCAUUACUUCAACUUUGUGCUAAUGACACUGGGCUACAAAG ............(((((((((((((..(((((((......--............)))))))..))).)))))((((...((((((...((....))...))))))...))))..))))). ( -38.07) >DroYak_CAF1 7152 118 + 1 CUGUUUUAUCGAUUUGGGGGCGGGCUCAAUGCCAUAAAAC--UGGGAUCUGAGUUGGCGCUCUGCCACGCCCCCAAUAUUCAUUACUUCAACUUUGUGCUAAUGACACUGGGCUGCAAAG ..........((.(((((((((((((((((.(((......--))).)).))))))(((.....))).)))))))))...))..........(((((((((..........))).)))))) ( -40.70) >DroAna_CAF1 6818 119 + 1 CCCUUUUAUUGAUUCG-GGGCGGGCACCAUGGCUUAAAAUAAAAGGGUCGGGCUUGGACCAUUGCCACGCCCUUAAAGUUCAUUAAGUUCAAAUUGUGUUAAUGACACUGUGCACCAAAU (((............)-))..((((((..(((((((((((..(((((.(((((..........))).)))))))...)))..))))).)))....(((((...))))).)))).)).... ( -30.30) >consensus CUGUUUUAUCGAUUUGUGGGCGGGCUCAAUGCCAUAAAAC__AAGGAUCUGAGUUGGCGCUUUGCCACGCCCCCAAUAUUCAUUACUUCAACUUUGUGCUAAUGACACUGGGCUACAAAG .................((((((((..(((((((....................)))))))..))).)))))...................(((((((((..........))).)))))) (-23.06 = -24.88 + 1.81)

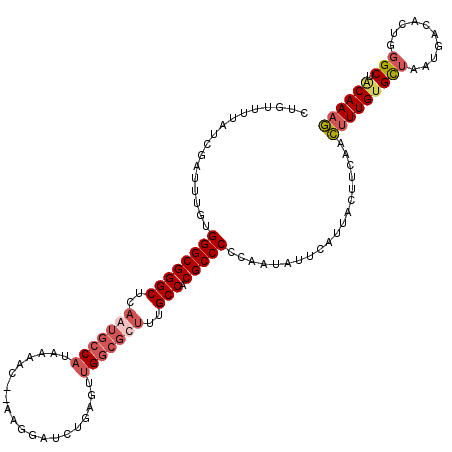
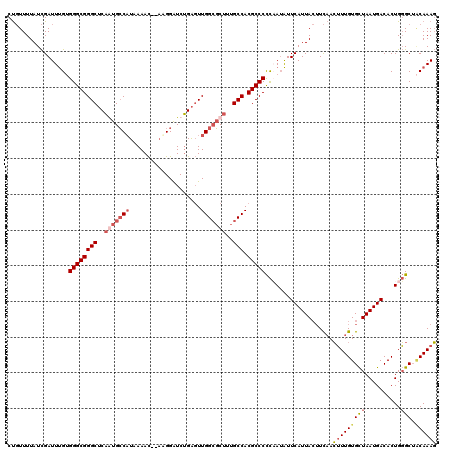
| Location | 10,354,425 – 10,354,543 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.49 |
| Mean single sequence MFE | -35.88 |
| Consensus MFE | -24.08 |
| Energy contribution | -24.77 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751985 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 10354425 118 - 22407834 CUUUGUAGCCCAGUGUCAUUAGCACAAAGUUGAAGUAAUGAAUAUGGGGGGCGUGGCAAAGCGCCAACUCAGAUCCUU--GUUUUAUGGCACUGAGCCCGCCCACAAAUCGAUAAAACAG .(((((..((((...((((((.((......))...))))))...))))(((((.(((..((.((((...(((....))--).....)))).))..)))))))))))))............ ( -38.50) >DroSim_CAF1 7125 118 - 1 CUUUGUAGCCCAGUGUCAUUAGCACAAAGUUGAAGUAAUGAAUAUGGGGGGCGUGGCAAAGCGCCAACUCAGAUCCAU--GUUUUAUGGCACUGAGCCCGCCCACAAAUCGAUAAAACAG .(((((..((((...((((((.((......))...))))))...))))(((((.(((..((.((((...((......)--).....)))).))..)))))))))))))............ ( -38.70) >DroYak_CAF1 7152 118 - 1 CUUUGCAGCCCAGUGUCAUUAGCACAAAGUUGAAGUAAUGAAUAUUGGGGGCGUGGCAGAGCGCCAACUCAGAUCCCA--GUUUUAUGGCAUUGAGCCCGCCCCCAAAUCGAUAAAACAG .....((((...((((.....))))...))))............(((((((((.(((.(((......))).(((.(((--......))).)))..))))))))))))............. ( -37.00) >DroAna_CAF1 6818 119 - 1 AUUUGGUGCACAGUGUCAUUAACACAAUUUGAACUUAAUGAACUUUAAGGGCGUGGCAAUGGUCCAAGCCCGACCCUUUUAUUUUAAGCCAUGGUGCCCGCCC-CGAAUCAAUAAAAGGG ....((.((((.((((.....))))........(((((........(((((((.(((..........))))).))))).....))))).....))))))..((-(............))) ( -29.32) >consensus CUUUGUAGCCCAGUGUCAUUAGCACAAAGUUGAAGUAAUGAAUAUGGGGGGCGUGGCAAAGCGCCAACUCAGAUCCUU__GUUUUAUGGCACUGAGCCCGCCCACAAAUCGAUAAAACAG .((((...((((...((((((..............))))))...))))(((((.(((..((.(((......................))).))..)))))))).))))............ (-24.08 = -24.77 + 0.69)


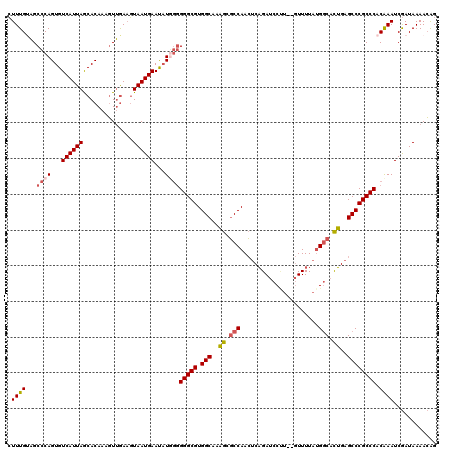
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:55:23 2006