
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 10,082,069 – 10,082,202 |
| Length | 133 |
| Max. P | 0.999200 |

| Location | 10,082,069 – 10,082,162 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 84.89 |
| Mean single sequence MFE | -21.76 |
| Consensus MFE | -16.60 |
| Energy contribution | -17.28 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.77 |
| SVM RNA-class probability | 0.976295 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
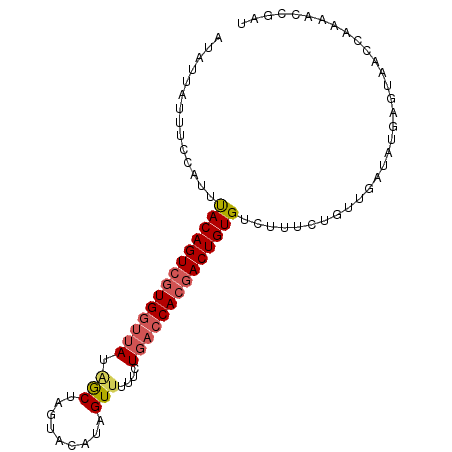
>2L_DroMel_CAF1 10082069 93 - 22407834 AUAUUAUUUCCACUUACAGUCGUGGCUAUGGCUUGU-CAUUGUUUUCCUGACCACGACUGUGUCUUUCUGUUGAUAUGAGUAACCAAAACCGAU ((((((........((((((((((((...(((....-....))).....).))))))))))).........))))))................. ( -20.83) >DroSec_CAF1 8796 94 - 1 AUAUUAUUUCCACUUACAGUCGUGGUUAUAACUAGCACAUUGUUUUUCUGACCACGACUGUGUCUUUCUGUUGAUAUGAGUAACCAAAACCAAU ((((((........((((((((((((((.((..(((.....))).)).)))))))))))))).........))))))................. ( -24.63) >DroSim_CAF1 8792 94 - 1 AUAUUAUUUCCAUUUACAGUCUUGGUUAUAACUAGCACAUAGUUUUUCUGACCACGACUGUGUCUUUCUGUUUAUUCUACUCACCAAAACCGAU ..............(((((((.((((((.(((((.....)))))....)))))).)))))))....((.((((.............)))).)). ( -19.12) >DroEre_CAF1 7590 94 - 1 AUACUAUUUCCAUUCACAGUGGUGCUGAUAGCUAGUAUAUCGUUUUUCUGACCACGACUGUGUCUUUCUAUUGAUUUGAGUAACCACAACCGAU .((((.........((((((.(((..((((.......))))(((.....)))))).))))))....((....))....))))............ ( -15.60) >DroYak_CAF1 8640 94 - 1 CUUUUAUUUCAAUUUACAGUCGUGGUUAUGGCUAGUAUAUAGUAUUCCUGACCACGACUGUGGCUUUCUGUUGAUAUGAGUAACCCAAACCGAU ..(((((.((((((((((((((((((((..((((.....)))).....)))))))))))))))......))))).))))).............. ( -28.60) >consensus AUAUUAUUUCCAUUUACAGUCGUGGUUAUAGCUAGUACAUAGUUUUUCUGACCACGACUGUGUCUUUCUGUUGAUAUGAGUAACCAAAACCGAU ..............((((((((((((((.(((.........)))....))))))))))))))................................ (-16.60 = -17.28 + 0.68)


| Location | 10,082,093 – 10,082,202 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 76.76 |
| Mean single sequence MFE | -23.75 |
| Consensus MFE | -14.03 |
| Energy contribution | -13.99 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982644 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
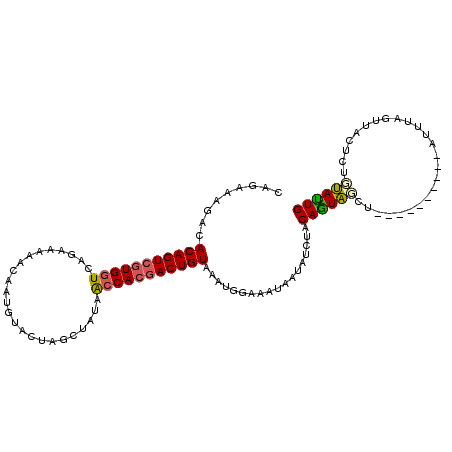
>2L_DroMel_CAF1 10082093 109 + 22407834 CAGAAAGACACAGUCGUGGUCAGGAAAACAAUG-ACAAGCCAUAGCCACGACUGUAAGUGGAAAUAAUAUCUACAGUAGCUUUAGUUACUUAUUUAGUUACUCUCUACUG .(((..((((((((((((((..((.........-.....))...)))))))))))..(((((.......)))))((((((....))))))......)))..)))...... ( -29.64) >DroSec_CAF1 8820 100 + 1 CAGAAAGACACAGUCGUGGUCAGAAAAACAAUGUGCUAGUUAUAACCACGACUGUAAGUGGAAAUAAUAUCUACAGUAGCU----------AUUUAGUUACUCUGUAUUG ((((.....(((((((((((.(....(((.........))).).)))))))))))..(((((.......))))).((((((----------....))))))))))..... ( -28.20) >DroSim_CAF1 8816 100 + 1 CAGAAAGACACAGUCGUGGUCAGAAAAACUAUGUGCUAGUUAUAACCAAGACUGUAAAUGGAAAUAAUAUCUACAGUAGCU----------AUUUAGUUACUCUGUAUUG ((((.....((((((.((((.(....(((((.....))))).).)))).))))))....................((((((----------....))))))))))..... ( -21.90) >DroEre_CAF1 7614 95 + 1 UAGAAAGACACAGUCGUGGUCAGAAAAACGAUAUACUAGCUAUCAGCACCACUGUGAAUGGAAAUAGUAUCCGCAAUGUA-------------AUACUC--AGUGCAUUG ........((((((.(((((.......))((((.......))))..))).))))))...(((.......))).(((((((-------------......--..))))))) ( -18.20) >DroYak_CAF1 8664 97 + 1 CAGAAAGCCACAGUCGUGGUCAGGAAUACUAUAUACUAGCCAUAACCACGACUGUAAAUUGAAAUAAAAGCUACAAUAUA-------------AUAUUCACACUGUAUUG .........(((((((((((..((....(((.....)))))...)))))))))))..................(((((((-------------..........))))))) ( -20.80) >consensus CAGAAAGACACAGUCGUGGUCAGAAAAACAAUGUACUAGCUAUAACCACGACUGUAAAUGGAAAUAAUAUCUACAGUAGCU__________AUUUAGUUACUCUGUAUUG .........(((((((((((........................)))))))))))..................((((((.........................)))))) (-14.03 = -13.99 + -0.04)


| Location | 10,082,093 – 10,082,202 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 76.76 |
| Mean single sequence MFE | -26.70 |
| Consensus MFE | -16.95 |
| Energy contribution | -17.43 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.27 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.63 |
| SVM decision value | 3.43 |
| SVM RNA-class probability | 0.999200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
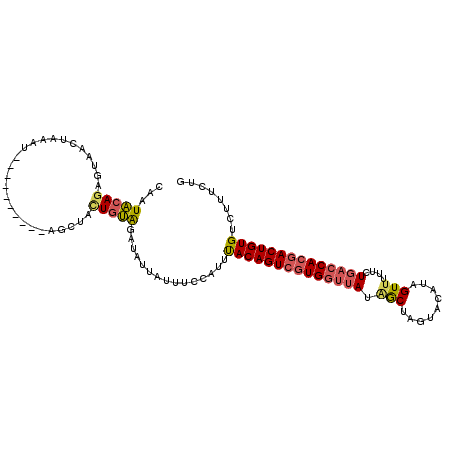
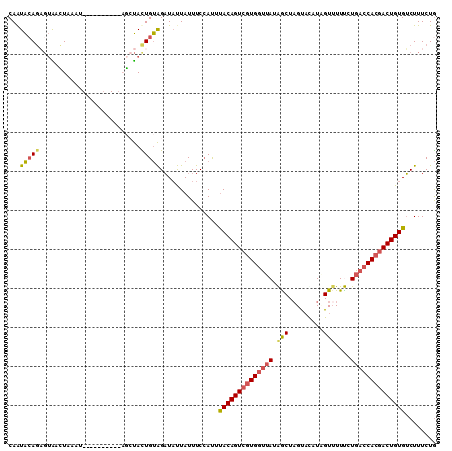
>2L_DroMel_CAF1 10082093 109 - 22407834 CAGUAGAGAGUAACUAAAUAAGUAACUAAAGCUACUGUAGAUAUUAUUUCCACUUACAGUCGUGGCUAUGGCUUGU-CAUUGUUUUCCUGACCACGACUGUGUCUUUCUG ((((((..(((.(((.....))).)))....)))))).................((((((((((((...(((....-....))).....).)))))))))))........ ( -29.40) >DroSec_CAF1 8820 100 - 1 CAAUACAGAGUAACUAAAU----------AGCUACUGUAGAUAUUAUUUCCACUUACAGUCGUGGUUAUAACUAGCACAUUGUUUUUCUGACCACGACUGUGUCUUUCUG ...((((((((........----------.))).)))))...............((((((((((((((.((..(((.....))).)).))))))))))))))........ ( -28.10) >DroSim_CAF1 8816 100 - 1 CAAUACAGAGUAACUAAAU----------AGCUACUGUAGAUAUUAUUUCCAUUUACAGUCUUGGUUAUAACUAGCACAUAGUUUUUCUGACCACGACUGUGUCUUUCUG ...((((((((........----------.))).)))))...............(((((((.((((((.(((((.....)))))....)))))).)))))))........ ( -24.70) >DroEre_CAF1 7614 95 - 1 CAAUGCACU--GAGUAU-------------UACAUUGCGGAUACUAUUUCCAUUCACAGUGGUGCUGAUAGCUAGUAUAUCGUUUUUCUGACCACGACUGUGUCUUUCUA ....(((.(--(.....-------------..)).)))(((.......)))...((((((.(((..((((.......))))(((.....)))))).))))))........ ( -20.00) >DroYak_CAF1 8664 97 - 1 CAAUACAGUGUGAAUAU-------------UAUAUUGUAGCUUUUAUUUCAAUUUACAGUCGUGGUUAUGGCUAGUAUAUAGUAUUCCUGACCACGACUGUGGCUUUCUG ...((((((((((...)-------------)))))))))..............(((((((((((((((..((((.....)))).....)))))))))))))))....... ( -31.30) >consensus CAAUACAGAGUAACUAAAU__________AGCUACUGUAGAUAUUAUUUCCAUUUACAGUCGUGGUUAUAGCUAGUACAUAGUUUUUCUGACCACGACUGUGUCUUUCUG ...(((((..........................)))))...............((((((((((((((.(((.........)))....))))))))))))))........ (-16.95 = -17.43 + 0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:53:10 2006