
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,957,169 – 9,957,261 |
| Length | 92 |
| Max. P | 0.902306 |

| Location | 9,957,169 – 9,957,261 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 71.82 |
| Mean single sequence MFE | -17.06 |
| Consensus MFE | -6.74 |
| Energy contribution | -6.46 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.40 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.767078 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9957169 92 + 22407834 AUUUAUGUCAAACGCUAACAUAAUCAUAAAACUAGAAAAUAACAGGGCAUCAUAUAACUGGUACACCUAUCUUCUAGUUGCUGUAGAUUUUU ..((((((.........))))))(((((.((((((((.((...((((.((((......)))).).))))).))))))))..))).))..... ( -13.90) >DroSec_CAF1 2606 92 + 1 AUUUAUGUCAACCGCUAACAACAUCAUAUAACUAGAAUAUGACAGGGUAUCCCACAUGGGGUACACCCAUCUUCUAGUUGCUGUAGUUUUGU .................((((.((.((((((((((((.(((...(((((((((....))))))).))))).))))))))).))).)).)))) ( -26.20) >DroSim_CAF1 2605 92 + 1 AUUUAUGUCAAACGCUAACAACAUCAUAUAACAAGAAUAUGACAGGGUAUCACACAGUCGGUACACCCAUCUUCUAGUUGCUGUAGUUUUAU .............((((.((((.((((((.......))))))..(((((((........))))).)).........))))...))))..... ( -15.00) >DroEre_CAF1 2619 81 + 1 AUUUAUUUUA-CUACUACAUUUACUAGUA----------UACCAGGGCAUCACAUAACCGGUGCACUAAUCAUCUAGUUGCUAUAGUUUUUU ..........-..((((.....(((((..----------....((.(((((........))))).))......))))).....))))..... ( -13.20) >DroYak_CAF1 2604 90 + 1 AUUUAUAUAA-CUACUUCAAUCACUAUAAAUUUAUAAAUUAACAGGGCAUCAAAUAGUCGGUGCACCAAUCAUCUAGUUGCAGUAGUUU-UU ........((-(((((.((((...((((....))))........(((((((........))))).)).........)))).))))))).-.. ( -17.00) >consensus AUUUAUGUCAACCGCUAACAUCAUCAUAAAACUAGAAAAUAACAGGGCAUCACAUAACCGGUACACCAAUCUUCUAGUUGCUGUAGUUUUUU ............................................(((((((........))))).))......................... ( -6.74 = -6.46 + -0.28)



| Location | 9,957,169 – 9,957,261 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 71.82 |
| Mean single sequence MFE | -20.54 |
| Consensus MFE | -8.24 |
| Energy contribution | -7.80 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.40 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902306 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
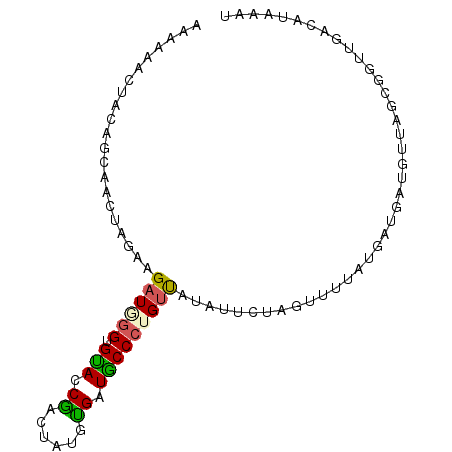
>2L_DroMel_CAF1 9957169 92 - 22407834 AAAAAUCUACAGCAACUAGAAGAUAGGUGUACCAGUUAUAUGAUGCCCUGUUAUUUUCUAGUUUUAUGAUUAUGUUAGCGUUUGACAUAAAU .........((..((((((((((((((.(((.((......)).)))))))))...))))))))...)).(((((((((...))))))))).. ( -22.50) >DroSec_CAF1 2606 92 - 1 ACAAAACUACAGCAACUAGAAGAUGGGUGUACCCCAUGUGGGAUACCCUGUCAUAUUCUAGUUAUAUGAUGUUGUUAGCGGUUGACAUAAAU .(((..(((((((((((((((((((((.(((.(((....))).)))))))))...))))))).......))))).)))...)))........ ( -27.11) >DroSim_CAF1 2605 92 - 1 AUAAAACUACAGCAACUAGAAGAUGGGUGUACCGACUGUGUGAUACCCUGUCAUAUUCUUGUUAUAUGAUGUUGUUAGCGUUUGACAUAAAU ...((((...((((((........((((((..((....))..)))))).(((((((.......)))))))))))))...))))......... ( -19.00) >DroEre_CAF1 2619 81 - 1 AAAAAACUAUAGCAACUAGAUGAUUAGUGCACCGGUUAUGUGAUGCCCUGGUA----------UACUAGUAAAUGUAGUAG-UAAAAUAAAU .....((((((...(((((...(((((.(((.((......)).))).))))).----------..)))))...))))))..-.......... ( -17.50) >DroYak_CAF1 2604 90 - 1 AA-AAACUACUGCAACUAGAUGAUUGGUGCACCGACUAUUUGAUGCCCUGUUAAUUUAUAAAUUUAUAGUGAUUGAAGUAG-UUAUAUAAAU ..-.((((((((((..((((((.((((....)))).)))))).))).((((.((((....)))).)))).......)))))-))........ ( -16.60) >consensus AAAAAACUACAGCAACUAGAAGAUGGGUGUACCGACUAUGUGAUGCCCUGUUAUAUUCUAGUUUUAUGAUGAUGUUAGCGGUUGACAUAAAU .....................((((((.(((.((......)).)))))))))........................................ ( -8.24 = -7.80 + -0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:22 2006