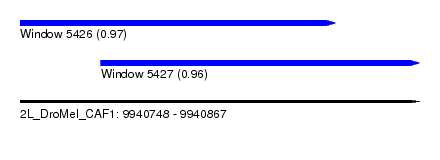

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,940,748 – 9,940,867 |
| Length | 119 |
| Max. P | 0.972098 |

| Location | 9,940,748 – 9,940,842 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 95.20 |
| Mean single sequence MFE | -18.67 |
| Consensus MFE | -17.02 |
| Energy contribution | -17.18 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.972098 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9940748 94 + 22407834 AAUAUGGGGCCAUAUCGGCCUUCUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU ..((..(((((.....)))))..))....((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... ( -20.20) >DroSec_CAF1 5464 94 + 1 AAUAUGGGGCCAUAUCGACCUUCUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU .......((((..............))))((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... ( -15.04) >DroSim_CAF1 5939 94 + 1 AAUAUGCGGCCAUAUCGGCCUUCUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU .......((((.....)))).........((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... ( -18.40) >DroEre_CAF1 6153 92 + 1 AA--UGAGGUCAUAUCGGCCUUUUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU ..--.((((((.....)))))).......((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... ( -17.40) >DroYak_CAF1 5710 92 + 1 AA--AGGGGCCGAAUCGGCCUUUUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU .(--((((((((...))))))))).....((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... ( -22.30) >consensus AAUAUGGGGCCAUAUCGGCCUUCUAGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAU .....((((((.....)))))).......((((((.....)))))).(((((((....((..((((((....))))))..)))))))))..... (-17.02 = -17.18 + 0.16)


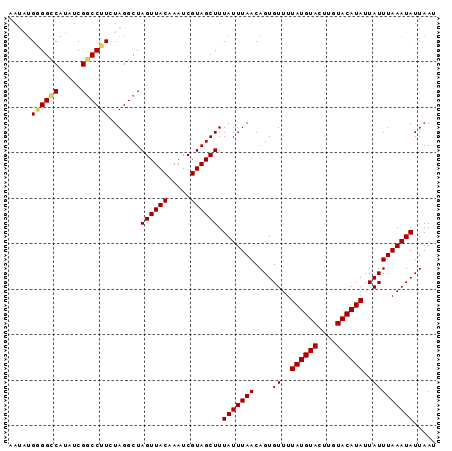
| Location | 9,940,772 – 9,940,867 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 99.37 |
| Mean single sequence MFE | -16.80 |
| Consensus MFE | -16.80 |
| Energy contribution | -16.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.64 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961060 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9940772 95 + 22407834 AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACAAU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... ( -16.80) >DroSec_CAF1 5488 95 + 1 AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACAAU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... ( -16.80) >DroSim_CAF1 5963 95 + 1 AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACAAU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... ( -16.80) >DroEre_CAF1 6175 95 + 1 AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACACU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... ( -16.80) >DroYak_CAF1 5732 95 + 1 AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACACU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... ( -16.80) >consensus AGGCUAGUUACAAAUCGUAGCUUUAUUUAACAGUGUUUUAUGUACUUGUACAUAUUAUUUAAAUAUUAAUAGCACUAAACAUAAUGGUCAACAAU .((((((((((.....)))))..........(((((..((((((....)))))).((((........)))))))))........)))))...... (-16.80 = -16.80 + 0.00)

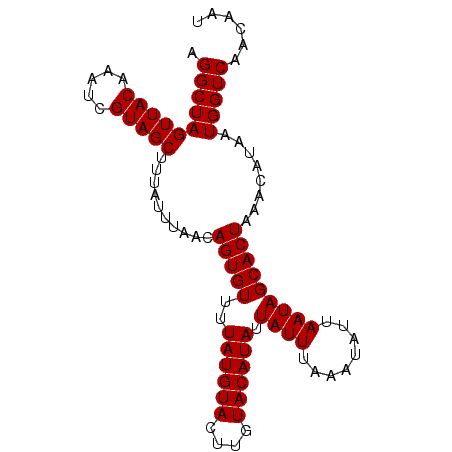

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:52:16 2006