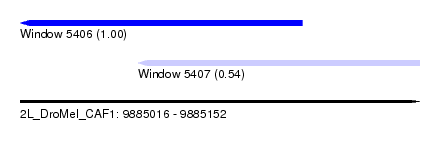

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,885,016 – 9,885,152 |
| Length | 136 |
| Max. P | 0.995813 |

| Location | 9,885,016 – 9,885,112 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 88.27 |
| Mean single sequence MFE | -26.19 |
| Consensus MFE | -21.45 |
| Energy contribution | -22.01 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.62 |
| SVM RNA-class probability | 0.995813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
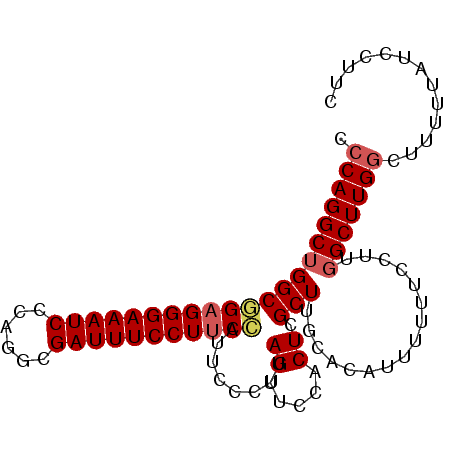
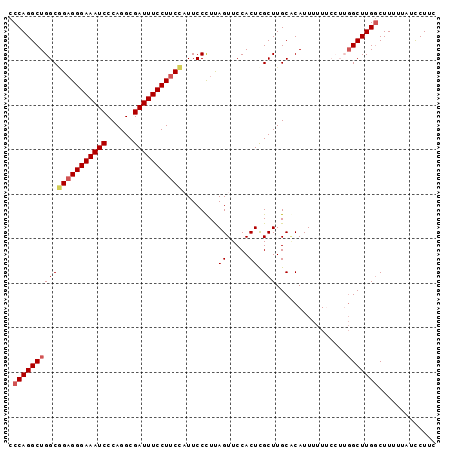
>2L_DroMel_CAF1 9885016 96 - 22407834 CCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACUCGCUUGCACAUUUUUUCCUUGGCUUGGCUUCUUAUCCUUC .((((((..(.(((((((((((......)))))))))))..................(....)...........)..))))))............. ( -26.90) >DroSec_CAF1 92934 96 - 1 CCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACUUGCUUGCGCAUUUUUUCCUUGGCUUGGCUUUUUAUCCUUC .((((((..(.(((((((((((......))))))))))).................(((....)))........)..))))))............. ( -29.70) >DroSim_CAF1 99788 96 - 1 CCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACUCGCUUGCGCAUUUUUUCCUUGGCUUGGCUUUUUAUCCUUC .((((((..(.(((((((((((......)))))))))))..................((....)).........)..))))))............. ( -29.30) >DroEre_CAF1 97841 91 - 1 CCCAGGCUGGCAGAGGGAAAUCCCAGGCGAUUUCCUUCUUUUCCUUCAGUUCCACUCGCUUGCCCAUUUUU-----GGCUUGGCUUUUUAUUCUGC .(((((((((((((((((((((......)))))))))).........(((.......))))))).......-----)))))))............. ( -25.61) >DroYak_CAF1 98940 91 - 1 CCCAGGCUGGCUGCGGGAAAUCCCAGGCGAUUUCCUCCUCUUCCUUUAGUUCCACUCGCUUGCACAUUUUU-----UGCUUGUCUUUCCAUUCUGC ....((..(((.(((((((((..(((((((.........((......))......)))))))...))))))-----)))..)))...))....... ( -19.46) >consensus CCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACUCGCUUGCACAUUUUUUCCUUGGCUUGGCUUUUUAUCCUUC .(((((((((((((((((((((......)))))))))))........((.....)).)))................)))))))............. (-21.45 = -22.01 + 0.56)

| Location | 9,885,056 – 9,885,152 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 95.62 |
| Mean single sequence MFE | -24.90 |
| Consensus MFE | -23.58 |
| Energy contribution | -23.94 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
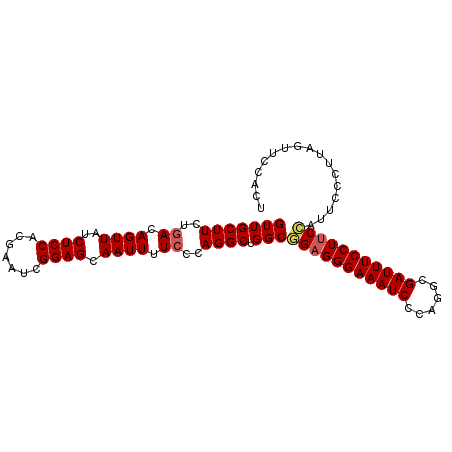
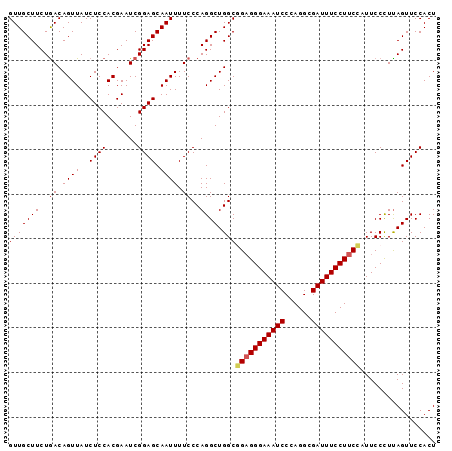
>2L_DroMel_CAF1 9885056 96 - 22407834 GUUGCUUCUAACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACU ((((....)))).................(((((.......((.....)).(((((((((((......))))))))))).........)))))... ( -25.60) >DroSec_CAF1 92974 96 - 1 GUUGCUUCUGACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACU (((((((..((.((((..((((.......)))).)))).))..)))).)))(((((((((((......)))))))))))................. ( -27.20) >DroSim_CAF1 99828 96 - 1 GUUGCUUCUGACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACU (((((((..((.((((..((((.......)))).)))).))..)))).)))(((((((((((......)))))))))))................. ( -27.20) >DroEre_CAF1 97876 96 - 1 GUUGCUUCUGACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCAGAGGGAAAUCCCAGGCGAUUUCCUUCUUUUCCUUCAGUUCCACU ((((((.((((..((.......))...)))))))))).......((((((.(((((((((((......)))))))))))......))))))..... ( -24.70) >DroYak_CAF1 98975 96 - 1 GUUGCUUCUGACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCUGCGGGAAAUCCCAGGCGAUUUCCUCCUCUUCCUUUAGUUCCACU ((((((.((((..((.......))...))))))))))......(((..((..(.((((((((......)))))))).).)).)))........... ( -19.80) >consensus GUUGCUUCUGACAGUUAUCUCCACGAAUCGGAGCAAUUUUCCCAGGCUGGCGGAGGGAAAUCCCAGGCGAUUUCCUUCCAUUCCCUUAGUUCCACU (((((((..((.((((..((((.......)))).)))).))..)))).)))(((((((((((......)))))))))))................. (-23.58 = -23.94 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:57 2006