
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,875,242 – 9,875,342 |
| Length | 100 |
| Max. P | 0.533115 |

| Location | 9,875,242 – 9,875,342 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 78.66 |
| Mean single sequence MFE | -32.00 |
| Consensus MFE | -20.83 |
| Energy contribution | -23.64 |
| Covariance contribution | 2.81 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.65 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.533115 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
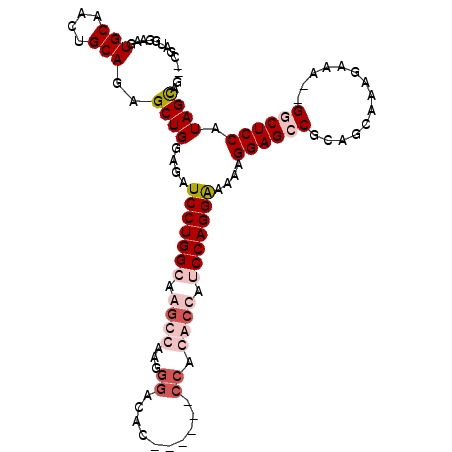
>2L_DroMel_CAF1 9875242 100 + 22407834 --CAAUGGAAGUGCAAUUGCAGAGCUGGAGAUCCUGGCAAGCCAAGGGACAC------CCAGGCUAUCCAGGAAAAAGGAGCCGCAGCAAAGAAA--GUCUCCAUAGCAG --..(((((..(((....)))..((((....((((((..((((..((....)------)..))))..))))))....(....).)))).......--...)))))..... ( -32.00) >DroSim_CAF1 90162 99 + 1 --CGAUGGAAGUGCAACUGCAGUGCUGGAGAUCCUGGCAAGCCAAGGGACAC------CC-GGCUAUCCAGGAAAAAGGAGCCGCAGCAAAGAAA--GGCUCCAUAGCAG --.........(((....))).(((((....((((((..((((..(((...)------))-))))..))))))....((((((.(......)...--)))))).))))). ( -37.50) >DroEre_CAF1 88335 106 + 1 --CGAUGGAACUGCAACGGCAGAGCUGGAGAUCCUGGGAAGCCGAGGGACCCUACAUGCCACACCAUCCAGGAAAAAGGAGCCGCACCAAAGAGA--GUCUCCAUAGUAG --.(((((..((((....)))).((.((.(.((((((....)).)))).))).....))....))))).........((((.(.(........).--).))))....... ( -26.60) >DroYak_CAF1 87569 100 + 1 GGCGAUGGAAGUGCAACUGCAGAGCUGGAGAUCCUGGG----------ACACCACAUCCCACAGCAUCCAGGGAAAAGGAGCCGCAGCAAAAAGAAGGGCUCCGUAGUGG .(((.......))).(((((....(((((.....((((----------(.......))))).....)))))......((((((.(..(.....)..)))))))))))).. ( -31.90) >consensus __CGAUGGAAGUGCAACUGCAGAGCUGGAGAUCCUGGCAAGCCAAGGGACAC______CCACACCAUCCAGGAAAAAGGAGCCGCAGCAAAGAAA__GGCUCCAUAGCAG ...........(((....)))..((((....(((((((.((((...((..........)).)))).)))))))....((((((..............)))))).)))).. (-20.83 = -23.64 + 2.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:47 2006