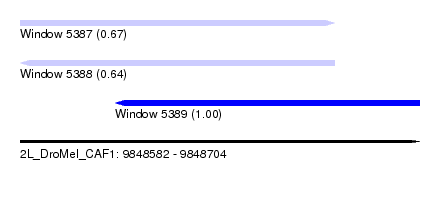
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,848,582 – 9,848,704 |
| Length | 122 |
| Max. P | 0.997241 |

| Location | 9,848,582 – 9,848,678 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 75.28 |
| Mean single sequence MFE | -23.27 |
| Consensus MFE | -18.43 |
| Energy contribution | -19.10 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.672311 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9848582 96 + 22407834 UGCGAGAUGCCUUCACUCCUUGGCGCCAACAAGGACGUCGCCAGGACCCAUCCAGGAAUCACACACAUAUACAUAUGUACUACACAUAUACGCACG ((((.(((.(((....((((.((((((.....))....)))))))).......))).)))............((((((.....)))))).)))).. ( -22.90) >DroSec_CAF1 57310 75 + 1 UGCGAGAUGCCUUUGGUCCUUGGCGCCAACGAGGACGACGCCAGGACCCAUCCAGGAUUCACACACA---------------------CACGCACU ((((.((((.....((((((.((((((.....))....))))))))))))))..(......).....---------------------..)))).. ( -22.70) >DroEre_CAF1 59888 73 + 1 UGCGAGAUGCCUUUGGUCCUUGGCGCCAAGGAGGACGUCGCCAGGACCCAUCCAGGAUCCACACACA---------------------C--GCACG ((((.((((((.....((((((....)))))))).))))....(((.((.....)).))).......---------------------)--))).. ( -24.20) >consensus UGCGAGAUGCCUUUGGUCCUUGGCGCCAACGAGGACGUCGCCAGGACCCAUCCAGGAUUCACACACA_____________________CACGCACG ((((.((((.....((((((.((((((.....))....))))))))))))))).(......).............................))).. (-18.43 = -19.10 + 0.67)

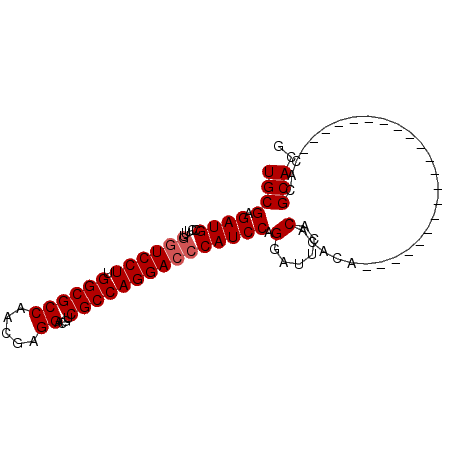
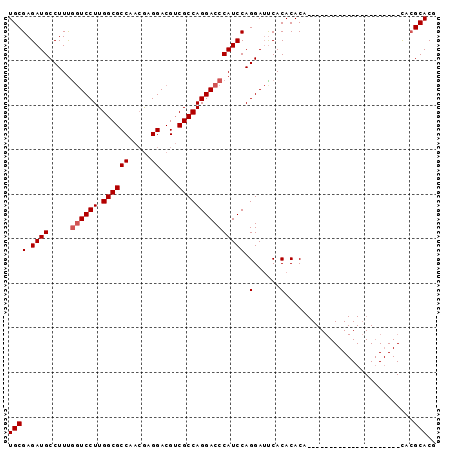
| Location | 9,848,582 – 9,848,678 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 75.28 |
| Mean single sequence MFE | -29.60 |
| Consensus MFE | -23.54 |
| Energy contribution | -23.43 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.637283 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9848582 96 - 22407834 CGUGCGUAUAUGUGUAGUACAUAUGUAUAUGUGUGUGAUUCCUGGAUGGGUCCUGGCGACGUCCUUGUUGGCGCCAAGGAGUGAAGGCAUCUCGCA ...((((((((((((.....))))))))))))(((.(((.(((...(.(.((((((((.((.......)).)))).)))).).)))).))).))). ( -30.70) >DroSec_CAF1 57310 75 - 1 AGUGCGUG---------------------UGUGUGUGAAUCCUGGAUGGGUCCUGGCGUCGUCCUCGUUGGCGCCAAGGACCAAAGGCAUCUCGCA ........---------------------..((((.((..(((.....(((((((((((((.......))))))).))))))..)))..)).)))) ( -30.40) >DroEre_CAF1 59888 73 - 1 CGUGC--G---------------------UGUGUGUGGAUCCUGGAUGGGUCCUGGCGACGUCCUCCUUGGCGCCAAGGACCAAAGGCAUCUCGCA .....--.---------------------..((((.(((((((.....((((((((((.((.......)).)))).))))))..))).)))))))) ( -27.70) >consensus CGUGCGUG_____________________UGUGUGUGAAUCCUGGAUGGGUCCUGGCGACGUCCUCGUUGGCGCCAAGGACCAAAGGCAUCUCGCA ...............................((((.((..(((.....((((((((((.((.......)).)))).))))))..)))..)).)))) (-23.54 = -23.43 + -0.11)

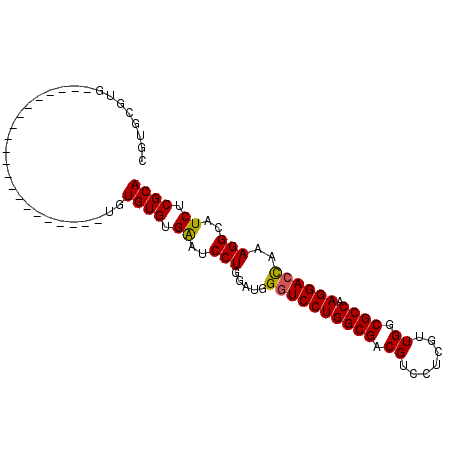
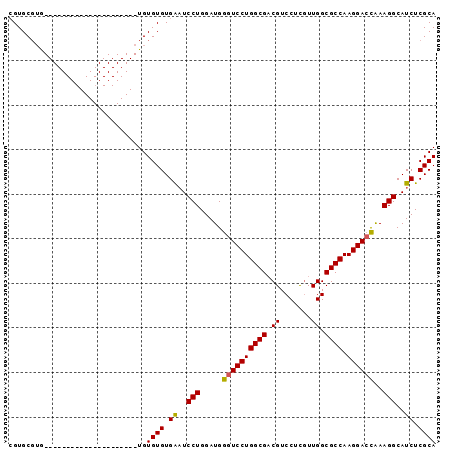
| Location | 9,848,611 – 9,848,704 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 71.71 |
| Mean single sequence MFE | -31.67 |
| Consensus MFE | -17.12 |
| Energy contribution | -17.56 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.54 |
| SVM decision value | 2.82 |
| SVM RNA-class probability | 0.997241 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9848611 93 - 22407834 CACACAAACAUACGCAUACUGGCGUGCGUGCGUAUAUGUGUAGUACAUAUGUAUAUGUGUGUGAUUCCUGGAUGGGUCCUGGCGACGUCCUUG ((((((....(((((((......))))))).(((((((((.....)))))))))...))))))......(((((.((....))..)))))... ( -30.40) >DroSec_CAF1 57339 70 - 1 CACACA--CAUACGCAUGCUGGCAUGAGUGCGUG---------------------UGUGUGUGAAUCCUGGAUGGGUCCUGGCGUCGUCCUCG ((((((--(((((((((.(......).)))))))---------------------))))))))......((((((((....)).))))))... ( -33.60) >DroEre_CAF1 59917 70 - 1 CACACACCCAUACGCAUGCAGGCUUGCGUGC--G---------------------UGUGUGUGGAUCCUGGAUGGGUCCUGGCGACGUCCUCC .......(((((((((((((.(....).)))--)---------------------))))))))).....(((((.((....))..)))))... ( -31.00) >consensus CACACA__CAUACGCAUGCUGGCAUGCGUGCGUG_____________________UGUGUGUGAAUCCUGGAUGGGUCCUGGCGACGUCCUCG ((((((....(((((((.(......).))))))).......................))))))......(((((.((....))..)))))... (-17.12 = -17.56 + 0.45)

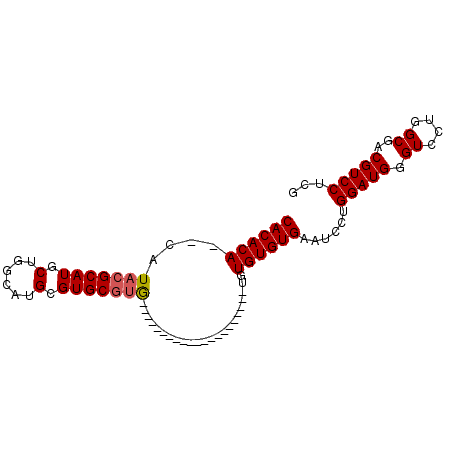
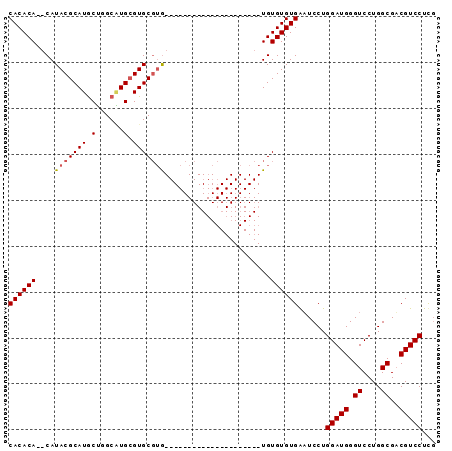
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:51:40 2006