
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,749,123 – 9,749,233 |
| Length | 110 |
| Max. P | 0.500000 |

| Location | 9,749,123 – 9,749,233 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.88 |
| Mean single sequence MFE | -33.48 |
| Consensus MFE | -19.83 |
| Energy contribution | -20.50 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.59 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
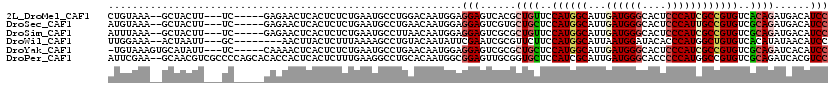
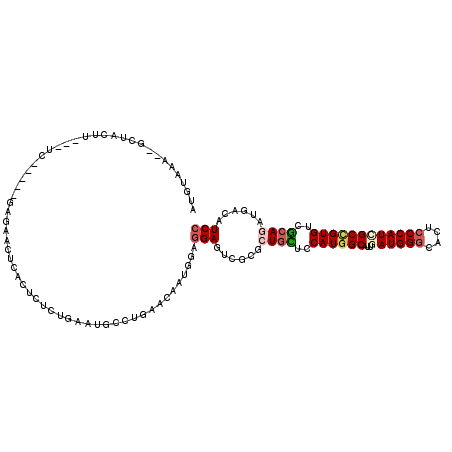
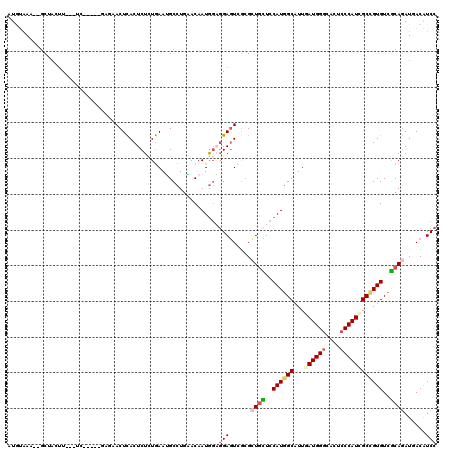
>2L_DroMel_CAF1 9749123 110 + 22407834 CUGUAAA--GCUACUU---UC-----GAGAACUCACUCUCUGAAUGCCUGGACAAUGGAGGAGUCACGCUGUUCCAUGGCAUUGAUGGGCACUCCCAUCGCCGUGUCACAGAUGACAUCC .(((...--((....(---((-----((((......)))).))).))....))).....(((((((..((((..((((((...((((((....))))))))))))..)))).)))).))) ( -36.60) >DroSec_CAF1 31254 110 + 1 AUGUAAA--GCUACUU---UC-----GAGAACUCACUCUCUGAAUGCCUGAACAAUGGAGGAGUCGUGCUGCUCCAUGGCAUUGAUGGGCACUCCCAUUGCCGUGUCGCAGAUGACAUCC .(((..(--((....(---((-----((((......)))).))).).))..))).....((((((((.((((..(((((((...(((((....))))))))))))..))))))))).))) ( -38.00) >DroSim_CAF1 30016 110 + 1 AUUUAAA--GCUACUU---UC-----GAGAACUCACUCUCUGAAUGCCUUAACAAUGGAGGAGUCGCGCUGUUCCAUGGCAUUGAUGGGCACUCCCAUCGCCGUGUCGCAGAUGACAUCC ..((((.--((....(---((-----((((......)))).))).)).)))).......(((((((..((((..((((((...((((((....))))))))))))..)))).)))).))) ( -36.00) >DroWil_CAF1 46546 107 + 1 UUGGAAA--ACUAAUU---GC--------AACUUACUCUUUAAAAGCCUGUACAAUAUUCGAAUCGCGUUCUUCCAUGGCAUUAAUGGAUACACCCAUGGCUGUGUCACAUAUAACAUCC .(((((.--......(---((--------(.(((.........)))..))))........((((...)))))))))((((((..((((......))))....))))))............ ( -15.10) >DroYak_CAF1 32461 111 + 1 -UGUAAAGUGCAUAUU---UC-----CAAAACUCACUCUCUGAAUGCCUGAACAAUGGAGGAGUCGCGCUGCUCCAUGGCAUUGAUGGGCACUCCCAUCGCCGUGUCGCAGAUCACAUCC -(((...(((.((.((---((-----((....(((..(.......)..)))....)))))).))))).((((..((((((...((((((....))))))))))))..))))...)))... ( -32.80) >DroPer_CAF1 31195 118 + 1 AUUCGAA--GCAACGUCGCCCCAGCACACCACUCACUCUUUGAAGGCCUGCACAAUGGCGGAGUUGCGGUGCUCCAUCGCAUUGAUGGGCACCCCCAUGGCCGUGUCGCAGAUCACGUCC .......--(((((..((((...(((..((..(((.....))).))..))).....))))..)))))(((((.(((((.....)))))))))).....((.((((........)))).)) ( -42.40) >consensus AUGUAAA__GCUACUU___UC_____GAGAACUCACUCUCUGAAUGCCUGAACAAUGGAGGAGUCGCGCUGCUCCAUGGCAUUGAUGGGCACUCCCAUCGCCGUGUCGCAGAUGACAUCC ...........................................................(((......((((..((((((...((((((....))))))))))))..))))......))) (-19.83 = -20.50 + 0.67)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:50:38 2006