
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,010,667 – 1,010,780 |
| Length | 113 |
| Max. P | 0.975037 |

| Location | 1,010,667 – 1,010,780 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.88 |
| Mean single sequence MFE | -27.49 |
| Consensus MFE | -21.68 |
| Energy contribution | -22.24 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.887775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1010667 113 + 22407834 UUAUUUGUAUUUGUAUAUUAUCAUCCGCGGCUACCAAACUCAAUCCUUCUGAAAGAAAACAGUGCGAAAACAGUG-GAUUCAAGUUUGGUCGCCUCUCCGAUAACUUUGUACGC------ ............((((((((((....(.(((.((((((((.(((((..(((...................))).)-))))..)))))))).))).)...)))))...)))))..------ ( -26.61) >DroSec_CAF1 122391 111 + 1 UUAUUUGUAUUUGUAUAUUAUCAUCCGCGGCCACCAAACUGAAUCCUUUUGAAAGAAGACAGUGCGU--UCAUUG-GAUUCAAGUUUGGUUGCCUCUCCGAUAACUUUGUACGC------ ............((((((((((....(.(((.((((((((((((((...((((....(......).)--)))..)-))))).)))))))).))).)...)))))...)))))..------ ( -32.00) >DroSim_CAF1 122953 110 + 1 UUAUUUGUAUUUGUAUAUUAUCAUCCGCGGCCACCAAACUGAAUCCUUUUGAAAGAAAACAGUGCGU--UCAUUG-GAUUCAAGU-UGGUUGCCUCUCCGAUAACUUUGUACGC------ ............((((((((((....(.(((.(((((..(((((((...((((.(.........).)--)))..)-))))))..)-)))).))).)...)))))...)))))..------ ( -28.90) >DroYak_CAF1 129635 118 + 1 UUAUUUGUAUUUGUAUAUUAUCAUCCGCGGUUACCAAACUGAAUCCUUUCGAAAAAAAACUGCUAGA--ACAUUGAGACUCAAGUUUGGUUGCCUCUCUGAUAACUUUGUACGCCAGACU ......((...((((((((((((...(.(((.((((((((......((((((...............--...))))))....)))))))).))).)..))))))...))))))....)). ( -22.47) >consensus UUAUUUGUAUUUGUAUAUUAUCAUCCGCGGCCACCAAACUGAAUCCUUUUGAAAGAAAACAGUGCGA__ACAUUG_GAUUCAAGUUUGGUUGCCUCUCCGAUAACUUUGUACGC______ ............((((((((((....(.(((.((((((((((((((............................).))))).)))))))).))).)...)))))...)))))........ (-21.68 = -22.24 + 0.56)
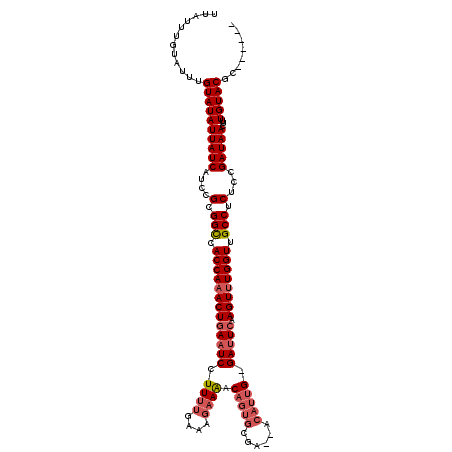


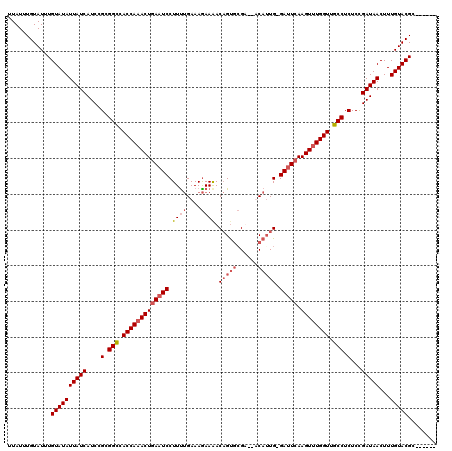
| Location | 1,010,667 – 1,010,780 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.88 |
| Mean single sequence MFE | -31.31 |
| Consensus MFE | -24.43 |
| Energy contribution | -24.81 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.975037 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
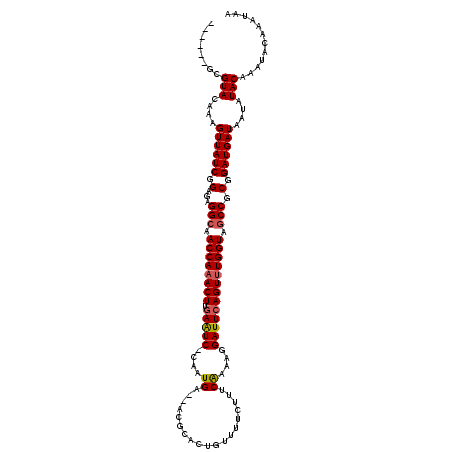
>2L_DroMel_CAF1 1010667 113 - 22407834 ------GCGUACAAAGUUAUCGGAGAGGCGACCAAACUUGAAUC-CACUGUUUUCGCACUGUUUUCUUUCAGAAGGAUUGAGUUUGGUAGCCGCGGAUGAUAAUAUACAAAUACAAAUAA ------..(((....((((((.(...(((.(((((((((.((((-(..(((....)))(((........)))..)))))))))))))).))).).))))))....)))............ ( -33.60) >DroSec_CAF1 122391 111 - 1 ------GCGUACAAAGUUAUCGGAGAGGCAACCAAACUUGAAUC-CAAUGA--ACGCACUGUCUUCUUUCAAAAGGAUUCAGUUUGGUGGCCGCGGAUGAUAAUAUACAAAUACAAAUAA ------..(((....((((((.(...(((.((((((((.(((((-(..(((--(.............))))...)))))))))))))).))).).))))))....)))............ ( -33.42) >DroSim_CAF1 122953 110 - 1 ------GCGUACAAAGUUAUCGGAGAGGCAACCA-ACUUGAAUC-CAAUGA--ACGCACUGUUUUCUUUCAAAAGGAUUCAGUUUGGUGGCCGCGGAUGAUAAUAUACAAAUACAAAUAA ------..(((....((((((.(...(((.((((-(.(((((((-(..(((--(.............))))...)))))))).))))).))).).))))))....)))............ ( -30.92) >DroYak_CAF1 129635 118 - 1 AGUCUGGCGUACAAAGUUAUCAGAGAGGCAACCAAACUUGAGUCUCAAUGU--UCUAGCAGUUUUUUUUCGAAAGGAUUCAGUUUGGUAACCGCGGAUGAUAAUAUACAAAUACAAAUAA ........(((....(((((((..(.((..((((((((.(((((....(((--....))).........(....))))))))))))))..)).)...))))))).)))............ ( -27.30) >consensus ______GCGUACAAAGUUAUCGGAGAGGCAACCAAACUUGAAUC_CAAUGA__ACGCACUGUUUUCUUUCAAAAGGAUUCAGUUUGGUAGCCGCGGAUGAUAAUAUACAAAUACAAAUAA ........(((....((((((.(...(((.((((((((.(((((....((...................))....))))))))))))).))).).))))))....)))............ (-24.43 = -24.81 + 0.38)

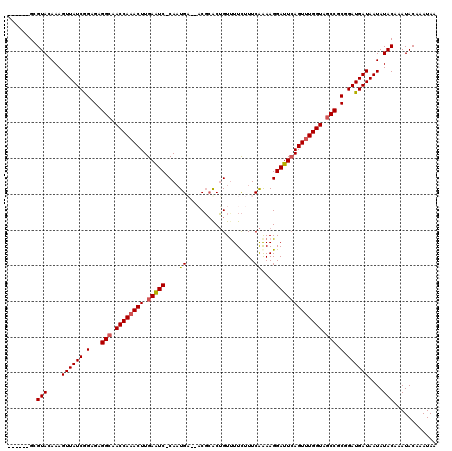
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:33:21 2006