| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,674,786 – 9,674,882 |
| Length | 96 |
| Max. P | 0.999483 |

| Location | 9,674,786 – 9,674,882 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 76.80 |
| Mean single sequence MFE | -23.24 |
| Consensus MFE | -18.42 |
| Energy contribution | -18.10 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.25 |
| Mean z-score | -3.49 |
| Structure conservation index | 0.79 |
| SVM decision value | 3.64 |
| SVM RNA-class probability | 0.999483 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9674786 96 + 22407834 AACCACCCAUUCAGACCACUCAGCCCACUCAGCCCGUUCAGCCAUUACGUUAUGGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCCACCCUU ......................((((((((.((..((((..((((......))))(((((.....))))))))).))....))))))))....... ( -27.00) >DroSec_CAF1 32931 77 + 1 AACCACCC------------------ACUCAGCCCACUCAACCAUUCCGUUCUGGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCCACCCU- ........------------------.....((((((((..(((........)))(((((.....)))))...........))))))))......- ( -25.10) >DroSim_CAF1 37741 77 + 1 AACCACCC------------------ACUCAACCCAUUCAGCCAUUCCGUUCUGGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCCACCCU- .....(((------------------((((...........(((........)))(((((.....)))))...........))))))).......- ( -21.30) >DroEre_CAF1 38624 72 + 1 AACC------------------ACCCACUCAGCCCACUCAGCCACUCCGUUCUAGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCC------ ....------------------.........((((((((....(((..((((...(((((.....))))))))))))....)))))))).------ ( -23.90) >DroYak_CAF1 33336 92 + 1 AACC---CACUCAACCCACUCAAUCCACUCAAUCCACUCAGCCAUUCCGUUCCAGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCCACCCA- ....---.......(((((((...............((((((......)))...)))(((.((((....)))).)))....))))))).......- ( -18.90) >consensus AACCACCC________________CCACUCAGCCCACUCAGCCAUUCCGUUCUGGAGGCUUUUCAAGCCUGAAUAGUAAUCGAGUGGGCCACCCU_ ...............................(((((((((((......)))....(((((.....)))))...........))))))))....... (-18.42 = -18.10 + -0.32)


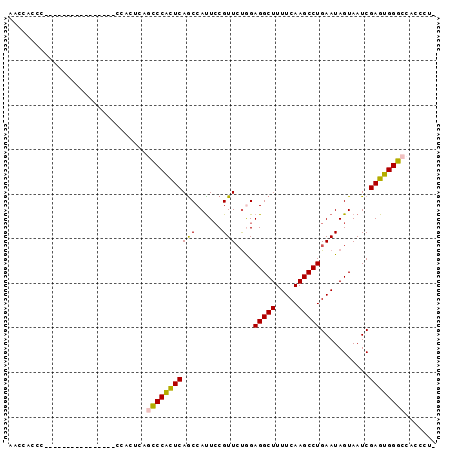
| Location | 9,674,786 – 9,674,882 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 76.80 |
| Mean single sequence MFE | -31.48 |
| Consensus MFE | -21.08 |
| Energy contribution | -21.68 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.10 |
| Structure conservation index | 0.67 |
| SVM decision value | 3.54 |
| SVM RNA-class probability | 0.999357 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
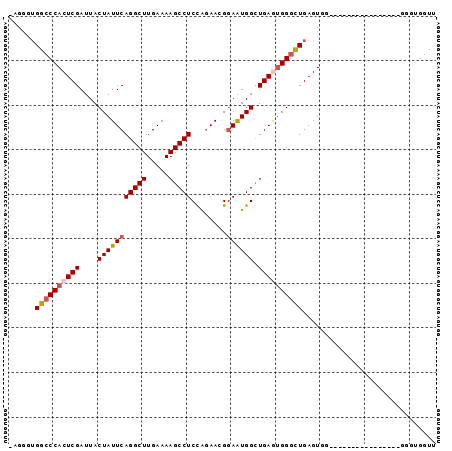
>2L_DroMel_CAF1 9674786 96 - 22407834 AAGGGUGGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCCAUAACGUAAUGGCUGAACGGGCUGAGUGGGCUGAGUGGUCUGAAUGGGUGGUU .....(.(((((.(((...(((((((((.((.....(((((((((......))))......)))))....)).))))))))).))).))))).).. ( -33.10) >DroSec_CAF1 32931 77 - 1 -AGGGUGGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCCAGAACGGAAUGGUUGAGUGGGCUGAGU------------------GGGUGGUU -....(.(((((((((....(((((((((((.....))))(((.....))).....)))))))..)))))------------------)))).).. ( -29.80) >DroSim_CAF1 37741 77 - 1 -AGGGUGGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCCAGAACGGAAUGGCUGAAUGGGUUGAGU------------------GGGUGGUU -....(.(((((((((((..(((((((((((.....))))(((.....))).....))))))))))))))------------------)))).).. ( -31.70) >DroEre_CAF1 38624 72 - 1 ------GGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCUAGAACGGAGUGGCUGAGUGGGCUGAGUGGGU------------------GGUU ------.(((((((((....(((((((((((.....)))((((.....))))...))))))))..)))))))))------------------.... ( -29.10) >DroYak_CAF1 33336 92 - 1 -UGGGUGGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCUGGAACGGAAUGGCUGAGUGGAUUGAGUGGAUUGAGUGGGUUGAGUG---GGUU -.....((((((((((((..((((((((.((.....((((((((...))))..))))....)).)))))))))))))))))))).....---.... ( -33.70) >consensus _AGGGUGGCCCACUCGAUUACUAUUCAGGCUUGAAAAGCCUCCAGAACGGAAUGGCUGAGUGGGCUGAGUGG________________GGGUGGUU ......((((((((((....(((((((((((.....)))))........)))))).)))))))))).............................. (-21.08 = -21.68 + 0.60)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:50:14 2006