
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,641,734 – 9,641,849 |
| Length | 115 |
| Max. P | 0.619212 |

| Location | 9,641,734 – 9,641,849 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.96 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -21.77 |
| Energy contribution | -21.00 |
| Covariance contribution | -0.77 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.619212 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
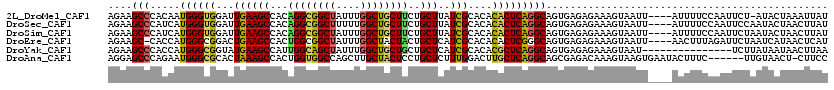
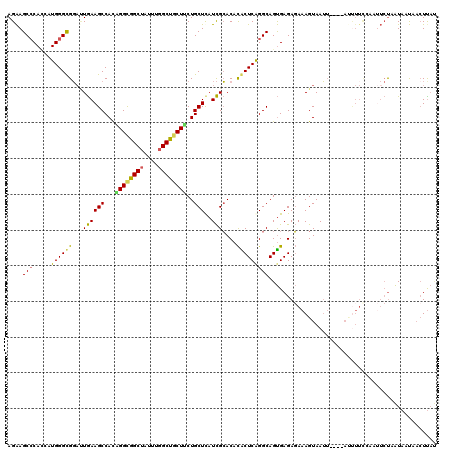
>2L_DroMel_CAF1 9641734 115 + 22407834 AGAAGCCCACAAUGGGUGGAUUGAAGCCACAGGCGGCUAUUUGGCUGCUUCUGCUUAUCGCACACACUCAGGCAGUGAGAGAAAGUAAUU----AUUUUCCAAUUCU-AUACUAAAUUAU ....((((.....))))((((((..(((..((((((((....)))))))).(((.....)))........))).......((((((....----)))))))))))).-............ ( -30.00) >DroSec_CAF1 42279 116 + 1 AGAAGCCCAUCAUGGGUGGAUUGAAGCCACAGGCGGCUUUUUGGCUGCUUCUGCUUAUCGCACACACUCAGGCAGUGAGAGAAAGUAAUU----AUUUUCCAAUUCCAAUACUAACUUAU ....((((.....))))((((((..(((..((((((((....)))))))).(((.....)))........))).......((((((....----)))))))))))).............. ( -29.90) >DroSim_CAF1 41741 116 + 1 AGAAGCCCAUCAUGGGUGGAUUGAAGCCACAGGCGGCUAUUUGGCUGCUUCUGCUUAUCGCACACACUCAGGCAGUGAGAGAAAGUAAUU----AUUUUCCAAUUCUAAUACUAACUUAU ....((((.....))))((((((..(((..((((((((....)))))))).(((.....)))........))).......((((((....----)))))))))))).............. ( -30.00) >DroEre_CAF1 44356 115 + 1 AGAAGC-CACCAUGGGCGGACUGAAGCCACUGGCGGCUAUUUGGCUACUACUGCUCAUCGCACACACUCGGGCAGUGAGAGAAAGUAAUU----AACUUUAGAUUCUAAUCAUAACUCAU ...(((-(.(((..(((........)))..))).))))..........((((((((.............)))))))).((((((((....----.))))).(((....)))....))).. ( -31.32) >DroYak_CAF1 44097 105 + 1 AGAAGCCCACCAUGGGCGGUAUGAAGCCAUUGGCAGCUAUUUGGCUGCUGCUGCUCAUCGCACACGCUCAGGCAGUGAGAGAAAGUAAU---------------UCUUAUAAUAACUUAA ....((((.....))))(((.....)))...(((((((....)))))))((((((....((....))...))))))..(((((.....)---------------))))............ ( -31.70) >DroAna_CAF1 54281 113 + 1 AGGAGCCCAGAAUGGGCGCACUAAAGCCACUGGUGGCCAGCUUGCUACUCCUGCUCUUUGGACUUGCUCAGGCAGCGAGACAAAGUAAGUGAAUACUUUC------UUGUAACU-CUUCC ((((((((.....))))(((.....((((....)))).....)))...))))((((((((..((((((.....)))))).)))))..)))...(((....------..)))...-..... ( -32.60) >consensus AGAAGCCCACCAUGGGCGGAUUGAAGCCACAGGCGGCUAUUUGGCUGCUUCUGCUCAUCGCACACACUCAGGCAGUGAGAGAAAGUAAUU____AUUUUCCAAUUCUAAUAAUAACUUAU ....(((.....((((((...((((((...((((((((....))))))))..)))..)))....)))))))))............................................... (-21.77 = -21.00 + -0.77)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:58 2006