
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,638,918 – 9,639,015 |
| Length | 97 |
| Max. P | 0.688646 |

| Location | 9,638,918 – 9,639,015 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.69 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -16.22 |
| Energy contribution | -16.05 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.688646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9638918 97 - 22407834 GGUAAAAAUGUAAUUUUCAAGUGAAGGUGAAGUUGCAGCC--GAGA------------------AACUGGCAG-AAAGUGAACUGCGAA-GGCUCUGGGGCCAA-AGGGCACAUGGAGAA ........((((((((.((........)))))))))).((--.(((------------------..((.((((-........))))..)-)..))).))(((..-..))).......... ( -23.50) >DroSec_CAF1 39494 97 - 1 GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGCC--GAGA------------------AACUGGCAG-AAAGUGAACUGCGAA-GGCUCUGUGGCCAA-AGGGCGCAUGGAGAA (((.....((((((((.((........)))))))))))))--....------------------..((.((((-........))))..)-).((((((((((..-..))).))))))).. ( -27.80) >DroSim_CAF1 38791 97 - 1 GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGCC--GAGA------------------AACUGGCAG-AAAGUGAACUGCGAA-GGCUCUGGGGCCAA-AGGGCGCAUGGAGAA ........((((((((.((........)))))))))).((--.(((------------------..((.((((-........))))..)-)..))).))(((..-..))).......... ( -25.50) >DroEre_CAF1 41660 97 - 1 GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGCA--CAGA------------------AACUGGCUG-AAAGUGAACUGGUAG-GGCUCUGGGGCCAA-AGGGCACAUGGAGAA .............((((((.(((..(((..(.((.((((.--(((.------------------..)))))))-.)).)..))).....-((((....))))..-....))).)))))). ( -23.80) >DroYak_CAF1 41250 103 - 1 GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGUUGCCAGUUU----------------CUCUGGCAG-AAAGUGAACUGGUAGGGGCUUUGGGGCCAAAAGGGCACAUGGCGAA (((((...((((((((.((........)))))))))).))))).....----------------(((((.(((-........))).)))))(((.((..(((.....))).)).)))... ( -32.80) >DroAna_CAF1 51483 116 - 1 GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGUU--GAGUUCCCUGGAACGAGAGCAGAAAUAGCAGCAAAGUGAACUCGGGA-GGCUCUAAGGUGAA-AGGGCAGGAAAAGAA ........(((.((((((....))))))....(..(....--((((..((((........(((....).)).....((....)))))).-.))))....)..).-...)))......... ( -22.10) >consensus GGUAAAAAUGCAAUUUUCAAGUGAAGGUGAAGUUGCAGCC__GAGA__________________AACUGGCAG_AAAGUGAACUGCGAA_GGCUCUGGGGCCAA_AGGGCACAUGGAGAA ........((((((((.((........))))))))))........................................(((..((......((((....))))...))..)))........ (-16.22 = -16.05 + -0.17)

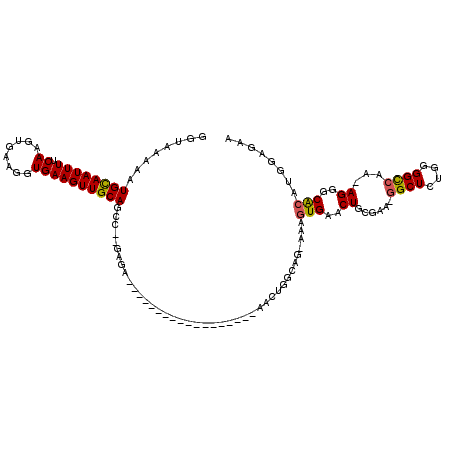
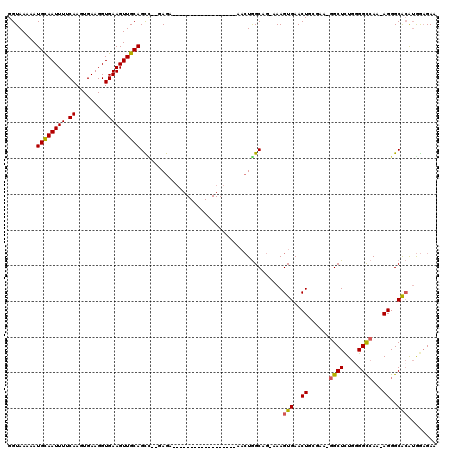
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:56 2006