| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,535,010 – 9,535,102 |
| Length | 92 |
| Max. P | 0.999438 |

| Location | 9,535,010 – 9,535,102 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 76.47 |
| Mean single sequence MFE | -33.64 |
| Consensus MFE | -22.67 |
| Energy contribution | -24.51 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.23 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.67 |
| SVM decision value | 3.60 |
| SVM RNA-class probability | 0.999438 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9535010 92 - 22407834 UGUCGACUGUGACACUUCACCGUCACGCUGUCGGCCACCGCUGAUGGUCCACCACCUCCAGGUCCACUGCCAACUGUCGACGGCAAUUGAUC .(((((..(((((........)))))(((((((((....((.(.(((.((..........)).)))).)).....)))))))))..))))). ( -31.30) >DroSec_CAF1 124 86 - 1 UGUCGACUGUGACACUUCACCGUCACGCUGUCGGCCACCAUUGGUGGUCCA---CCUCCUGGGCC---GCCAACUGCCGACGGCAAUUGAUC .(((((..(((((........)))))(((((((((.....(((((((((((---.....))))))---)))))..)))))))))..))))). ( -44.20) >DroSim_CAF1 124 86 - 1 UGUCGACUGUGGCAUUUCACCGUCACGCUGUCGGUCACCACUGCUGGUCCA---CCUCCAGGGCC---GCUAACUGCCGACGGCAACUGAUC .((((((.((((....)))).)))..(((((((((.......((.(((((.---......)))))---)).....)))))))))....))). ( -33.50) >DroEre_CAF1 134 86 - 1 UGUCGACUGUGGCACUUCACCGUCACGCCGUCGGCCACCACUGGUGGUCCA---CCACCUGCGCC---CCCAAUUGCCGACGGCAACUGAUC .((((((.((((....)))).)))..(((((((((.......(((((....---)))))((....---..))...)))))))))....))). ( -34.50) >DroAna_CAF1 139 71 - 1 UGCCGACUGUGGCACUUGACCAUCACACUGUCCGGCC-CA-------CCCA---CUGCCUGGGCC----------GACGACGCCACCGUAUC .....((.(((((...((.....))....((((((((-((-------....---.....))))))----------)..)))))))).))... ( -24.70) >consensus UGUCGACUGUGGCACUUCACCGUCACGCUGUCGGCCACCACUGGUGGUCCA___CCUCCUGGGCC___GCCAACUGCCGACGGCAACUGAUC ....(((.((((....)))).)))..(((((((((..........((((((........))))))..........)))))))))........ (-22.67 = -24.51 + 1.84)

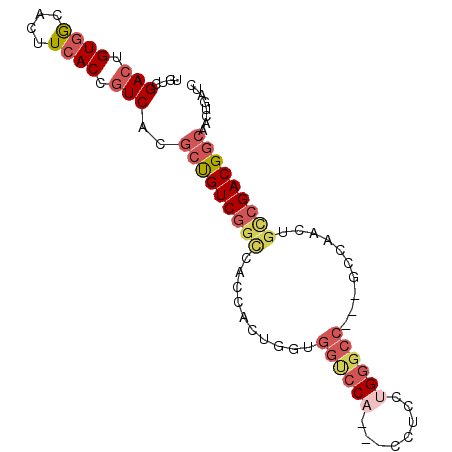

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:08 2006