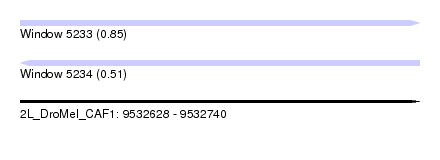

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,532,628 – 9,532,740 |
| Length | 112 |
| Max. P | 0.850863 |

| Location | 9,532,628 – 9,532,740 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 87.68 |
| Mean single sequence MFE | -36.12 |
| Consensus MFE | -27.51 |
| Energy contribution | -28.22 |
| Covariance contribution | 0.71 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.850863 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9532628 112 + 22407834 AAGUGAGCAACUGUUACUGCUCUCACCCGGAUGGUUAUUGCGAUGACGA------CGAUGGCGACGCCAUCG-AAUCUUUUUUGCAUCUUGACGAGGGCAUAGCAGUUUCAUUAGCCCU .(((((..((((((((.(((((((..(.((((((((((....)))))..------(((((((...)))))))-...........))))).)..))))))))))))))))))))...... ( -41.00) >DroSec_CAF1 40014 110 + 1 AAGUGAGCAACUGUUACUGCUCUCACCCGGAUGGUUAUUGCGAUGA---------CGAUGGCGACGCCAUCGGAAUCUUUUUUGCAUCUUGACGAGGACAUAGCAGUUUCAUUAGCCCU .(((((..((((((((.((..(((..(.(((((((....))(((..---------(((((((...)))))))..))).......))))).)..)))..)))))))))))))))...... ( -37.00) >DroSim_CAF1 43162 98 + 1 AAGUGAGCAACUGUUACUACUCUCACCCGGAUGGUUA---------------------UGGCGACGCCAUCGGAAUCUUUUUUGCAUCUUGACGAGGGCAUAGCAGUUUCAUUAGCCCU .(((((..((((((((...(((((..(.(((((...(---------------------((((...)))))(((((....)))))))))).)..)))))..)))))))))))))...... ( -26.50) >DroEre_CAF1 43904 118 + 1 AAGUGAGCAACUGUUACUGCUCUCACCCGGAUGGUUAUUGUGAUGAUGACGAUGACGAUGACGACGCCAUUGGAAUCUU-UUUGCAUCUUGACGAGGGCAUAGCAGUUUCAUUAGCCCU .(((((..((((((((.(((((((..(.(((((((((((.....))))))(((..(((((.......)))))..)))..-....))))).)..))))))))))))))))))))...... ( -39.30) >DroYak_CAF1 47978 119 + 1 AAGUGAGCAACUGUUACUGCUCUCACCCGGAUGGUUAUUGCGAUGACGAUGAUGACGACGACGACGCCAUUGGAAUCUUUUUUGCAUCUUGACGAGGGCAUAGCAGUUUCAUUAACCCU .(((((..((((((((.(((((((..(.(((((((....))(((..(((((.((.((....)).)).)))))..))).......))))).)..))))))))))))))))))))...... ( -36.80) >consensus AAGUGAGCAACUGUUACUGCUCUCACCCGGAUGGUUAUUGCGAUGA_GA______CGAUGGCGACGCCAUCGGAAUCUUUUUUGCAUCUUGACGAGGGCAUAGCAGUUUCAUUAGCCCU .(((((..((((((((.(((((((..(.((((((((((((...............)))))))((..((...))..)).......))))).)..))))))))))))))))))))...... (-27.51 = -28.22 + 0.71)


| Location | 9,532,628 – 9,532,740 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 87.68 |
| Mean single sequence MFE | -32.21 |
| Consensus MFE | -24.02 |
| Energy contribution | -23.34 |
| Covariance contribution | -0.68 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.75 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.514962 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9532628 112 - 22407834 AGGGCUAAUGAAACUGCUAUGCCCUCGUCAAGAUGCAAAAAAGAUU-CGAUGGCGUCGCCAUCG------UCGUCAUCGCAAUAACCAUCCGGGUGAGAGCAGUAACAGUUGCUCACUU .((((...((..((((((..((((((.....))..........(((-((((((((.((....))------.)))))))).)))........))))...))))))..))...)))).... ( -36.10) >DroSec_CAF1 40014 110 - 1 AGGGCUAAUGAAACUGCUAUGUCCUCGUCAAGAUGCAAAAAAGAUUCCGAUGGCGUCGCCAUCG---------UCAUCGCAAUAACCAUCCGGGUGAGAGCAGUAACAGUUGCUCACUU .((((...((..((((((.....(((..(..((((.......(((..(((((((...)))))))---------..)))........))))..)..)))))))))..))...)))).... ( -36.96) >DroSim_CAF1 43162 98 - 1 AGGGCUAAUGAAACUGCUAUGCCCUCGUCAAGAUGCAAAAAAGAUUCCGAUGGCGUCGCCA---------------------UAACCAUCCGGGUGAGAGUAGUAACAGUUGCUCACUU .((((...((..((((((..((((((.....)).........(((....(((((...))))---------------------)....))).))))...))))))..))...)))).... ( -26.60) >DroEre_CAF1 43904 118 - 1 AGGGCUAAUGAAACUGCUAUGCCCUCGUCAAGAUGCAAA-AAGAUUCCAAUGGCGUCGUCAUCGUCAUCGUCAUCAUCACAAUAACCAUCCGGGUGAGAGCAGUAACAGUUGCUCACUU .((((...((..((((((..((((((.....))......-..(((..(.((((((.......)))))).)..)))................))))...))))))..))...)))).... ( -31.10) >DroYak_CAF1 47978 119 - 1 AGGGUUAAUGAAACUGCUAUGCCCUCGUCAAGAUGCAAAAAAGAUUCCAAUGGCGUCGUCGUCGUCAUCAUCGUCAUCGCAAUAACCAUCCGGGUGAGAGCAGUAACAGUUGCUCACUU .((((...((..((((((..((((((.....))(((......(((....((((((....))))))....)))......)))..........))))...))))))..))...)))).... ( -30.30) >consensus AGGGCUAAUGAAACUGCUAUGCCCUCGUCAAGAUGCAAAAAAGAUUCCGAUGGCGUCGCCAUCG______UC_UCAUCGCAAUAACCAUCCGGGUGAGAGCAGUAACAGUUGCUCACUU .((((...((..((((((..((((((.....))................(((((...))))).............................))))...))))))..))...)))).... (-24.02 = -23.34 + -0.68)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:49:07 2006