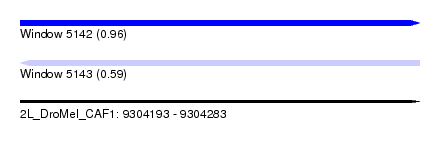

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 9,304,193 – 9,304,283 |
| Length | 90 |
| Max. P | 0.960065 |

| Location | 9,304,193 – 9,304,283 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 90.38 |
| Mean single sequence MFE | -22.48 |
| Consensus MFE | -17.70 |
| Energy contribution | -18.78 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.960065 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9304193 90 + 22407834 AAAAAUAAAAAUGAAUGGCCAACGAACAUGACCAGAGGCGGUGCGUUUAAAAUAAUUGUUAGCAAAGGGGCUCUGUUG------GCCACAAA----AAAU ...............(((((((((........)((((.(..(((...(((.....)))...)))....).))))))))------))))....----.... ( -20.40) >DroSec_CAF1 133667 93 + 1 A------AAAAUGAAUGGCCAACGAACAUGACCAGAGGCGGUGCGUUUAAAAUAAUUGUUAGCAAAGGGGCUCUGUUGGCGGUGGCCACAAAAA-AAAAU .------........((((((.((....((((.((((.(..(((...(((.....)))...)))....).)))))))).)).))))))......-..... ( -22.80) >DroSim_CAF1 136757 100 + 1 AAAAAUAAAAAUGAAUGGCCAACGAACAUGACCAGAGGCGGUGCGUUUAAAAUAAUUGUUAGCAAAGGGGCUCUGUUGGCGGUGGCCACAAAAAAAAAAU ...............((((((.((....((((.((((.(..(((...(((.....)))...)))....).)))))))).)).))))))............ ( -22.80) >DroEre_CAF1 144139 97 + 1 AAAAAUAAAAAUGAAUGGCCAACGAACAUGACCAGAGGCGGUACGUUUAAAAUAAUUGUUAGCAAAGGGGCUCUGUUGGCGGUGGCCACGAA---AAAAU ...............(((((...((((...(((......)))..))))......((((((((((.((...)).)))))))))))))))....---..... ( -21.80) >DroYak_CAF1 142117 96 + 1 AAAAAUAAAAAUGAAUGGCC-ACGAAUAUGACCAGAGGCAGUGCGUUUAAAAUAAUUGUUGGCAAAGGGGCUCUGUUGGUGGUGGCCACAAA---AAAAU ...............(((((-((.(...((((.((((.(..(((...(((.....)))...)))....).)))))))).).)))))))....---..... ( -24.60) >consensus AAAAAUAAAAAUGAAUGGCCAACGAACAUGACCAGAGGCGGUGCGUUUAAAAUAAUUGUUAGCAAAGGGGCUCUGUUGGCGGUGGCCACAAA___AAAAU ...............((((((.((....((((.((((.(..(((...(((.....)))...)))....).)))))))).)).))))))............ (-17.70 = -18.78 + 1.08)
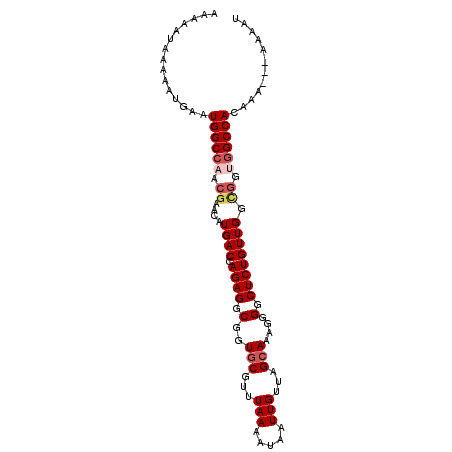


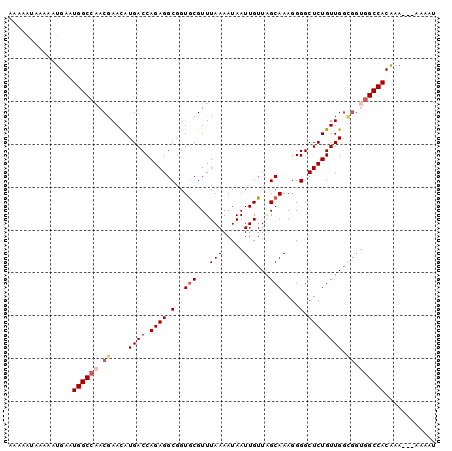
| Location | 9,304,193 – 9,304,283 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 90.38 |
| Mean single sequence MFE | -18.76 |
| Consensus MFE | -13.23 |
| Energy contribution | -14.31 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.587510 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 9304193 90 - 22407834 AUUU----UUUGUGGC------CAACAGAGCCCCUUUGCUAACAAUUAUUUUAAACGCACCGCCUCUGGUCAUGUUCGUUGGCCAUUCAUUUUUAUUUUU ....----...(((((------((((.((((....(((....)))...........(.((((....)))))..))))))))))))).............. ( -19.70) >DroSec_CAF1 133667 93 - 1 AUUUU-UUUUUGUGGCCACCGCCAACAGAGCCCCUUUGCUAACAAUUAUUUUAAACGCACCGCCUCUGGUCAUGUUCGUUGGCCAUUCAUUUU------U .....-.....(((((((.((..((((.(((......)))................(.((((....))))).)))))).))))))).......------. ( -18.20) >DroSim_CAF1 136757 100 - 1 AUUUUUUUUUUGUGGCCACCGCCAACAGAGCCCCUUUGCUAACAAUUAUUUUAAACGCACCGCCUCUGGUCAUGUUCGUUGGCCAUUCAUUUUUAUUUUU ...........(((((((.((..((((.(((......)))................(.((((....))))).)))))).))))))).............. ( -18.20) >DroEre_CAF1 144139 97 - 1 AUUUU---UUCGUGGCCACCGCCAACAGAGCCCCUUUGCUAACAAUUAUUUUAAACGUACCGCCUCUGGUCAUGUUCGUUGGCCAUUCAUUUUUAUUUUU .....---...(((((((.((.......(((......))).............(((((((((....)))).))))))).))))))).............. ( -19.00) >DroYak_CAF1 142117 96 - 1 AUUUU---UUUGUGGCCACCACCAACAGAGCCCCUUUGCCAACAAUUAUUUUAAACGCACUGCCUCUGGUCAUAUUCGU-GGCCAUUCAUUUUUAUUUUU .....---...((((((((......(((((...(..(((.................)))..).))))).........))-)))))).............. ( -18.69) >consensus AUUUU___UUUGUGGCCACCGCCAACAGAGCCCCUUUGCUAACAAUUAUUUUAAACGCACCGCCUCUGGUCAUGUUCGUUGGCCAUUCAUUUUUAUUUUU ...........(((((((..((.((((..(((.....((......................))....)))..)))).))))))))).............. (-13.23 = -14.31 + 1.08)

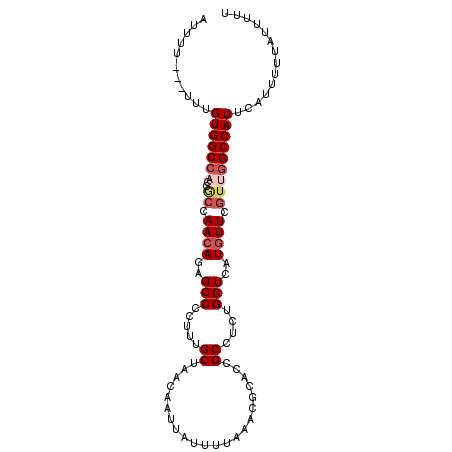
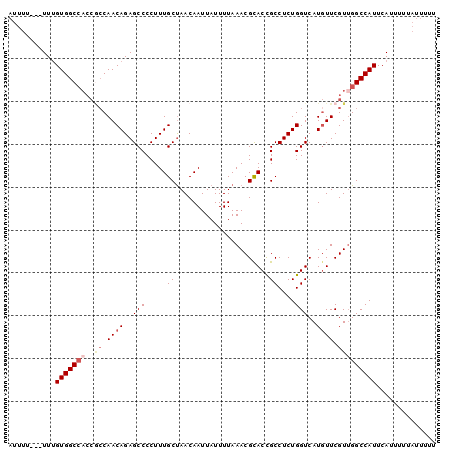
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:47:41 2006